[Job] Staff Associate II (Computational Structural Biology) at Columbia University
The X3DNA-DSSR resource is at the forefront of structural bioinformatics, developing advanced tools for analyzing and modeling nucleic acid structures. We are seeking a highly motivated Staff Associate II to join our team and contribute to our next-generation analysis and visualization engine.
To see our resource in action, please visit wDSSR, our new web interface for dissecting and modeling 3D nucleic acid structures: https://web.x3dna-dssr.org/.
We are looking for a candidate with a strong scientific background in structural biology or bioinformatics and a desire to contribute to peer-reviewed publications through community-driven data analysis. We value individuals who are eager to learn, adapt to new technical challenges, and support the global research community.
For the full job description and to submit your application, please visit the official Columbia University posting:
https://apply.interfolio.com/183705
Announcing wDSSR: The Next-Generation Web Interface to X3DNA-DSSR
Dear 3DNA/DSSR Community,
We are thrilled to announce the official launch of wDSSR (https://web.x3dna-dssr.org/), the powerful new web interface to the X3DNA-DSSR analytical engine.
Developed by Drs. Shuxiang Li and Xiang-Jun Lu and supported by NIH grant R24GM153869, wDSSR represents a major leap forward from our highly popular 2019 Web 3DNA 2.0 framework. While Web 3DNA 2.0 has faithfully served the community for the analysis, visualization, and modeling of 3D nucleic acid structures, wDSSR was built from the ground up to take full advantage of modern web technologies and the latest DSSR backend capabilities.
A Modern, Streamlined Scientific Workflow
We have completely overhauled the user interface to provide a clean, intuitive, and task-driven experience. The core modeling and analysis tools are now seamlessly organized into a logical, single-word scientific workflow: Analyze, Rebuild, Model, Circularize, Mutate, Assemble, and Visualize.
Spotlight Feature: The "Assemble" Module
One of the most exciting upgrades is the newly renamed Assemble tab (formerly "Composite"). This advanced composite model builder allows you to effortlessly construct complex, higher-order models by linking any combination of nucleic acid duplexes or protein-DNA/RNA complexes. You can quickly connect up to six distinct target structures, ranging from simple linked A-DNA and B-DNA duplexes to large, protein-decorated structural assemblies.
Immediate Global Adoption
Although wDSSR has just launched, we are incredibly humbled to share that it is already seeing rapid worldwide adoption! According to recent network infrastructure data, the new interface is actively being used by researchers across North America, South America, Europe, and Asia. Within just a few days, we have recorded active sessions from prestigious institutions around the globe, including:
- The Weizmann Institute of Science in Israel
- Katholieke Universiteit Leuven in Belgium
- Queen's University in Canada
- Universidad Nacional Autonoma de Mexico (UNAM) in Mexico
- Emory University and the Wadsworth Centers Laboratories and Research in the United States
- Jawaharlal Nehru University and the China Education and Research Network in Asia
How to Cite
While a dedicated paper for wDSSR is currently in preparation, researchers should cite the server using its URL (https://web.x3dna-dssr.org/) alongside the 2019 Web 3DNA 2.0 paper and the foundational 2015 DSSR paper. Full details and funding acknowledgements can be found on our newly consolidated About page.
We invite you all to try out the new wDSSR platform! As always, your feedback is invaluable to us, and we encourage you to share your thoughts, questions, and structural models via the newly updated Questions & Feedback link in the wDSSR footer.
Happy modeling!
Over the past few weeks, I’ve had the pleasure to talk to Thomas Holder, the PyMOL Principal Developer at Schrödinger, on possible integration of DSSR into PyMOL. On Tuesday April 21, 2015, I wrote to Thomas:
Last year, I had the please to collaborate with Dr. Robert Hanson to integrate DSSR into Jmol, see
http://chemapps.stolaf.edu/jmol/jsmol/dssr.htm. I am wondering if you have any interest in connecting DSSR to PyMOL. This will not only benefit both parties, but also bring elaborate analyses of RNA structures to the general audience. As you may be aware, RNA is becoming increasing important, yet the field of RNA structural bioinformatics is lagging (far) behind that of proteins.
After a few meet-ups, we all agree that the DSSR-PyMOL integration project would be meaningful/significant for RNA structural bioinformatics. Moreover, the community not only can benefit from the end result, but also should be able to make direct contributions through the process. On Friday May 08, 2015, Thomas sent out the following open invitation, titled Someone interested in writing a DSSR plugin for PyMOL?, to the PyMOL mailing list:
Is anyone interested in writing a DSSR plugin for PyMOL? DSSR is an integrated software tool for Dissecting the Spatial Structure of RNA (http://x3dna.bio.columbia.edu/docs/dssr-manual.pdf). Among other things, DSSR defines the secondary structure of RNA from 3D atomic coordinates in a way similar to DSSP does for proteins. Most of its output could be translated 1:1 into PyMOL selections, making it available for coloring and other selection based features. A PyMOL plugin could act as a wrapper which runs DSSR for an object or atom selection. Xiang-Jun Lu, the author of DSSR, is also working on base pair visualization (see http://x3dna.org/articles/seeing-is-understanding-as-well-as-believing), similar to (but more advanced) what’s already available from 3DNA (http://pymolwiki.org/index.php/3DNA).
Xiang-Jun would be happy to collaborate with someone who has experience with Python and the PyMOL API for writing an extension or plugin. Please contact me if this sounds appealing to you.
Get DSSR from http://x3dna.org/
See it hooked up with JSmol: http://chemapps.stolaf.edu/jmol/jsmol/dssr.htm
If you are self-motivated, care about software quality, have expertise in writing PyMOL plugin, and feel the pain in RNA structural analysis/visualization with currently available tools, now it is the time to make a difference. The DSSR/PyMOL project would ideally be composed of a team of dedicated practitioners with complementary skills. We will communicate mostly via email or online forum, in a presumably open and highly interactive way. By working on the project, you will be able to sharpen your skills and make new friends. The end product would not only make RNA structural bioinformatics easier for yourself but also benefit the community at large.

The v1.2.1 (2015feb01) release of DSSR contains a new functionality to characterize the so-called H-type pseudoknots. In this classical and most common type of pseudoknots, nucleotides from a hairpin loop form Watson-Crick base pairs with a single-stranded region outside of the hairpin to create another (adjacent) stem, as shown in the following illustration (taken from the Huang et al. paper A heuristic approach for detecting RNA H-type pseudoknots).

Normally, L2 is absent (i.e., with zero nucleotides) due to direct coaxial stacking of the two stems. An example output of DSSR on 1ymo (a human telomerase RNA pseudoknot) is shown below:

The corresponding sections from DSSR output are:
****************************************************************************
List of 3 H-type pseudoknot loop segments
1 stem#1(hairpin#1) vs stem#2(hairpin#2) L1 groove=MAJOR nts=8 UUUUUCUC U7,U8,U9,U10,U11,C12,U13,C14
2 stem#1(hairpin#1) vs stem#2(hairpin#2) L2 groove=----- nts=0
3 stem#1(hairpin#1) vs stem#2(hairpin#2) L3 groove=minor nts=8 CAAACAAA C30,A31,A32,A33,C34,A35,A36,A37
****************************************************************************
Secondary structures in dot-bracket notation (dbn) as a whole and per chain
>1ymo-1-A #1 nts=47 [chain] RNA
GGGCUGUUUUUCUCGCUGACUUUCAGCCCCAAACAAAAAAGUCAGCA
[[[[[[........(((((((((]]]]]]........))))))))).
Checking against the three-dimensional image and the secondary structure in linear form shown above, the meaning of the new section should be obvious. If you want to see more details, click the link to the DSSR-output file on 1ymo.

Recently I came across the following two citations to DSSR:
Base pair types were annotated with RNAview (45,46). Hydrogen bonds were annotated manually and with the help of DSSR of the 3DNA package (47,48). Helix parameters were obtained using the Curves+ web server (49). Structural figures were prepared using PyMol (50).
It is interesting to note that DSSR is cited here for its identification of hydrogen bonds, not its annotation of base pairs, among many other features. The simple geometry-based H-bonding identification algorithm, originally implemented in find_pair/analyze of 3DNA (and adopted by RNAView) and highly refined in DSSR, works well for nucleic acid structures. With the --get-hbonds option, users can now use DSSR as a tool just for its list of H-bonds outside of the program.
All figures were generated using PyMOL (60) or Chimera (48). The secondary structure diagram of the human mitoribosomal RNA was prepared by extracting base pairs from the model using DSSR (61). The secondary structure diagram was drawn in VARNA (62) and finalized in Inkscape.
I am very pleased to see that DSSR was cited for its ‘intended’ use in this important piece of work from a leading laboratory in structural biology. In the middle of last November (2013), I was approached by the lead author for proper citation of DSSR, and I suggested the two 3DNA papers. As far as I can remember, this was the first time I received such a question on DSSR citation. It prompted to write a FAQ entry in the DSSR User Manual, titled “How to cite DSSR?”. Hopefully, this citation issue will be gone in the near future.
Over the past two years, I’ve devoted significant efforts to make DSSR a handy tool for RNA structural bioinformatics; it certainly represents my view as to what a scientific software program should be like. As time passes by, DSSR is becoming increasingly sophisticated and citations to DSSR can only be higher.

Recently, PDB begins to release atomic coordinates of large (ribosomal) structures in mmCIF format. For nucleic-acid-containing structures, the largest one so far is 4v4g, the crystal structure of five 70S ribosomes from Escherichia coli in complex with protein Y. It is assembled from ten PDB entries (1voq, 1vor, 1vos, 1vou, 1vov, 1vow, 1vox, 1voy, 1voz, 1vp0), consisting of 22,345 nucleotides, and a total of 717,805 atoms.
This humongous structure poses no problems to DSSR at all, as shown below.
Command: x3dna-dssr -i=4v4g.cif -o=4v4g.out
Processing file '4v4g.cif' [4v4g]
total number of base pairs: 9277
total number of multiplets: 918
total number of helices: 1099
total number of stems: 1221
total number of isolated WC/wobble pairs: 603
total number of atom-base stacking interactions: 1736
total number of hairpin loops: 504
total number of bulges: 170
total number of internal loops: 775
total number of junctions: 214
total number of non-loop single-stranded segments: 429
total number of kissing loops: 5
total number of A-minor (type I and II) motifs: 100
total number of ribose zippers: 58 (1159)
total number of kink turns: 39
Time used: 00:00:10:45
It took less than 11 minutes to run on an iMac (and nearly 14 minutes on a Ubuntu Linux machine). Given the

From early on, 3DNA and DSSR have native support of modified nucleotides. The currently distributed baselist.dat file with 3DNA contains over 700 entries. As of v1.1.4-2014aug09, a new section has been added to DSSR to list explicitly the modified nucleotides in an analyzed structure.
Using the 76-nucleotide long yeast phenylalanine tRNA (1ehz) as an example, the pertinent section in DSSR output is as below.
List of 11 types of 14 modified nucleotides
nt count list
1 1MA-a 1 A.1MA58
2 2MG-g 1 A.2MG10
3 5MC-c 2 A.5MC40,A.5MC49
4 5MU-t 1 A.5MU54
5 7MG-g 1 A.7MG46
6 H2U-u 2 A.H2U16,A.H2U17
7 M2G-g 1 A.M2G26
8 OMC-c 1 A.OMC32
9 OMG-g 1 A.OMG34
10 PSU-P 2 A.PSU39,A.PSU55
11 YYG-g 1 A.YYG37
So 1ehz has 14 modified nucleotides of 11 different type, as listed in the following rows after the header line. The meaning of each column should be obvious. For example, the third row means that 5MC (5-methylcytidine, abbreviated as 'c' in 1-letter code) occurs twice, identified as A.5MC40 and A.5MC49, respectively.
With the 3-letter id, one can search the RCSB ligand database for more information about a specified modified nucleotide. The URL would be like this, using pseudouridine (PSU) as an example, https://www.rcsb.org/ligand/PSU.
It is hoped that the newly added section, put at the very top of DSSR output, will draw more attention to modified nucleotides.

From v1.1.3-2014jun18, DSSR has an additional output of RNA secondary structures in BPSEQ format. A sample file for PDB entry 1msy is shown below.
![1msy [GUAA tetra loop] in 3d and 2d representations 1msy [GUAA tetra loop] in 3d and 2d representations](http://forum.x3dna.org/images/1msy-3d-2d.png)
Filename: dssr-2ndstrs.bpseq
Organism: DSSR-derived secondary structure [1msy]
Accession Number: DSSR v1.1.4-2014aug09 (xiangjun@x3dna.org)
Citation: Please cite 3DNA/DSSR (see http://x3dna.org)
1 U 0 # name=A.U2647
2 G 26 # name=A.G2648, pairedNt=A.U2672
3 C 25 # name=A.C2649, pairedNt=A.G2671
4 U 24 # name=A.U2650, pairedNt=A.A2670
5 C 23 # name=A.C2651, pairedNt=A.G2669
6 C 22 # name=A.C2652, pairedNt=A.G2668
7 U 0 # name=A.U2653
8 A 0 # name=A.A2654
9 G 0 # name=A.G2655
10 U 0 # name=A.U2656
11 A 0 # name=A.A2657
12 C 17 # name=A.C2658, pairedNt=A.G2663
13 G 0 # name=A.G2659
14 U 0 # name=A.U2660
15 A 0 # name=A.A2661
16 A 0 # name=A.A2662
17 G 12 # name=A.G2663, pairedNt=A.C2658
18 G 0 # name=A.G2664
19 A 0 # name=A.A2665
20 C 0 # name=A.C2666
21 C 0 # name=A.C2667
22 G 6 # name=A.G2668, pairedNt=A.C2652
23 G 5 # name=A.G2669, pairedNt=A.C2651
24 A 4 # name=A.A2670, pairedNt=A.U2650
25 G 3 # name=A.G2671, pairedNt=A.C2649
26 U 2 # name=A.U2672, pairedNt=A.G2648
27 G 0 # name=A.G2673
Based on online sources, BPSEQ has originated from the Comparative RNA Web site developed by the Gutell lab. CRW files contain four header lines, describing the file name, organism, accession number, and a general remark. Thereafter, there is one line per base in the molecule, listing the position of the base (starting from 1), the one-letter base name (A,C,G,U etc), and the position number of the base to which it is paired. If the base is unpaired, zero (0) is put in the third column. In the above sample BPSEQ file derived from DSSR, detailed information about the base and its paired base (if any) comes after the # symbol.
Compared to dot-bracket notation (dbn) and connect-table (.ct) format, BPSEQ is simpler but less expressive. Nevertheless, the format is well-supported in bioinformatic tools on RNA secondary structures. It only seems fitting that DSSR now produces secondary structures in .bpseq (with default file name dssr-2ndstrs.bpseq), in addition to .dbn and .ct. Technically, adding the BPSEQ output to DSSR is trivial given the infrastructure already in place.

From early on, DSSR-derived RNA secondary structures in dot-bracket notation (dbn) have taken pseudoknots into consideration. Nevertheless, in DSSR releases prior to v1.1.3-2014jun18, the dbn output had been simplified to the first level only, with matched []s, even for RNA structures with high-order pseudoknots. RNA pseudoknot is a (relatively) complicated issue, and I’d planned to put off the topic until DSSR is well-established.
In early May, I noticed the Antczak et al. article RNApdbee—a webserver to derive secondary structures from pdb files of knotted and unknotted RNAs. I was delighted to read the following citation:
In order to facilitate a more comprehensive study, the webserver integrates the functionality of RNAView, MC-Annotate and 3DNA/DSSR, being the most common tools used for automated identification and classification of RNA base pairs.
Even before any paper on DSSR has been published, the software has already be ranked in the top three for the identification and classification of RNA base pairs! Well familiar with RNAView and MC-Annotate, I am glad to see DSSR is now listed on a par with them. Note that DSSR has far more functionality than just identifying and classifying RNA base pairs.
Further down the RNApdbee paper, especially in Figure 2, I found the following remarks regarding DSSR’s capability on RNA structures with high-order pseudoknot.
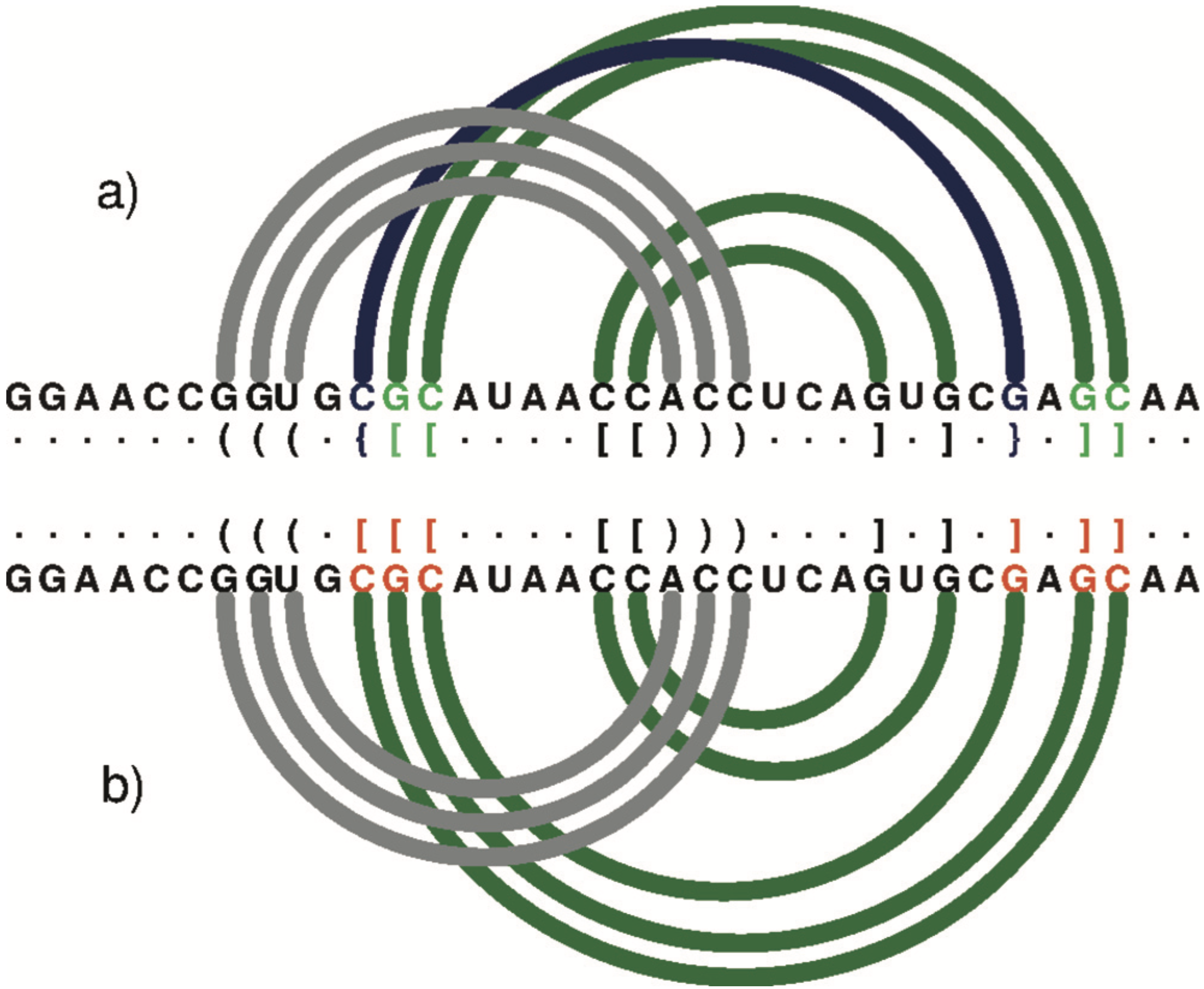
An arc diagram to represent the secondary structure of 1DDY (chain A) generated by R-CHIE upon the dot-bracket notation. Arcs of the same colour define a paired region. Crossing arcs reflect a conflict observed between the corresponding regions. (a) RNApdbee recognizes pseudoknots of the first (dark green) and second (navy blue) order. (b) 3DNA/DSSR improperly classifies base pairs (within residues in red) and the structure is recognized as the first-order pseudoknot.
The above citation and the question Higher-order pseudoknots in DP output (from Jan Hajic, Charles University in Prague) on the 3DNA Forum prompted me to further refine DSSR’s algorithm for deriving secondary structures of RNA with high-order pseudoknots. The DSSR v1.1.3-2014jun18 release made this revised functionality explicit. For the above cited PDB entry 1ddy, the relevant output of running DSSR on it would be:
Running command: "x3dna-dssr -i=1ddy.pdb"
****************************************************************************
This structure contains 2-order pseudoknot(s)
****************************************************************************
Secondary structures in dot-bracket notation (dbn) as a whole and per chain
>1ddy nts=140 [whole]
GGAACCGGUGCGCAUAACCACCUCAGUGCGAGCAA&GGAACCGGUGCGCAUAACCACCUCAGUGCGAGCAA&GGAACCGGUGCGCAUAACCACCUCAGUGCGAGCAA&GGAACCGGUGCGCAUAACCACCUCAGUGCGAGCAA
......(((.{[[....[[)))...].].}.]]..&......(((.{[[....[[)))...].].}.]]..&......(((.{[[....[[)))...].].}.]]..&......(((.{[[....[[)))...].].}.]]..
>1ddy-A #1 nts=35 [chain] RNA
GGAACCGGUGCGCAUAACCACCUCAGUGCGAGCAA
......(((.{[[....[[)))...].].}.]]..
>1ddy-C #2 nts=35 [chain] RNA
GGAACCGGUGCGCAUAACCACCUCAGUGCGAGCAA
......(((.{[[....[[)))...].].}.]]..
>1ddy-E #3 nts=35 [chain] RNA
GGAACCGGUGCGCAUAACCACCUCAGUGCGAGCAA
......(((.{[[....[[)))...].].}.]]..
>1ddy-G #4 nts=35 [chain] RNA
GGAACCGGUGCGCAUAACCACCUCAGUGCGAGCAA
......(((.{[[....[[)))...].].}.]]..
Note that the whole 1ddy entry contains four RNA chains (A, C, E, and G), and DSSR can handle each properly. So at least from DSSR v1.1.3-2014jun18, the following statement is no longer valid:
3DNA/DSSR improperly classifies base pairs (within residues in red) and the structure is recognized as the first-order pseudoknot.
A closely related issue is knot removal, a topic nicely summarized by Smit et al. in their publication From knotted to nested RNA structures: A variety of computational methods for pseudoknot removal. While not explicitly documented, the --nested (abbreviated to --nest) option has been available since DSSR v1.1.3-2014jun18. This option was first mentioned in the release note of DSSR v1.1.4-2014aug09. Again, using PDB entry 1ddy as an example, the relevant output of running DSSR with option --nested is as follows:
Running command: "x3dna-dssr -i=1ddy.pdb --nested"
****************************************************************************
This structure contains 2-order pseudoknot(s)
o You've chosen to remove pseudo-knots, leaving only nested pairs
****************************************************************************
Secondary structures in dot-bracket notation (dbn) as a whole and per chain
>1ddy nts=140 [whole]
GGAACCGGUGCGCAUAACCACCUCAGUGCGAGCAA&GGAACCGGUGCGCAUAACCACCUCAGUGCGAGCAA&GGAACCGGUGCGCAUAACCACCUCAGUGCGAGCAA&GGAACCGGUGCGCAUAACCACCUCAGUGCGAGCAA
......(((..........))).............&......(((..........))).............&......(((..........))).............&......(((..........))).............
>1ddy-A #1 nts=35 [chain] RNA
GGAACCGGUGCGCAUAACCACCUCAGUGCGAGCAA
......(((..........))).............
>1ddy-C #2 nts=35 [chain] RNA
GGAACCGGUGCGCAUAACCACCUCAGUGCGAGCAA
......(((..........))).............
>1ddy-E #3 nts=35 [chain] RNA
GGAACCGGUGCGCAUAACCACCUCAGUGCGAGCAA
......(((..........))).............
>1ddy-G #4 nts=35 [chain] RNA
GGAACCGGUGCGCAUAACCACCUCAGUGCGAGCAA
......(((..........))).............

H-bonding interactions are crucial for defining RNA secondary and tertiary structures. DSSR/3DNA contains a geometrically based algorithm for identifying H-bonds in nucleic-acid or protein structures given in .pdb or .cif format. Over the years, the method has been continuously refined, and it has served its purpose quite well. As of v1.1.1-2014apr11, this functionality is directly available from DSSR thorough the --get-hbonds option.
The output for 1msy, which contains a GUAA tetraloop mutant of Sarcin/Ricin domain from E. Coli 23 S rRNA, is listed below. The first line gives the header (# H-bonds in '1msy.pdb' identified by DSSR ...). The second line provides the total number of H-bonds (40) identified in the structure. Afterwards, each line consists of 8 space-delimited columns used to characterize a specific H-bond. Using the first one (#1) as an example, the meaning of each of the 8 columns is:
- The serial number (15), as denoted in the .pdb or .cif file, of the first atom of the H-bond.
- The serial number (578) of the second H-bond atom.
- The H-bond index (#1), from 1 to the total number of H-bonds.
- A one-letter symbol showing the atom-pair type (p) of the H-bond. It is ‘p’ for a donor-acceptor atom pair; ‘o’ for a donor/acceptor (such as the 2′-hydorxyl oxygen) with any other atom; ‘x’ for a donor-donor or acceptor-acceptor pair (as in #17); ‘?’ if the donor/acceptor status is unknown for any H-bond atom.
- Distance in Å between donor/acceptor atoms (2.768).
- Elemental symbols of the two atoms involved in the H-bond (O/N).
- Identifier of the first H-bond atom (O4@A.U2647).
- Identifier of the second H-bond atom (N1@A.G2673).
Command: x3dna-dssr -i=1msy.pdb --get-hbonds –o=1msy-hbonds.txt
# H-bonds in '1msy.pdb' identified by 3DNA version 3 (xiangjun@x3dna.org)
40
15 578 #1 p 2.768 O:N O4@A.U2647 N1@A.G2673
35 555 #2 p 2.776 O:N O6@A.G2648 N3@A.U2672
36 554 #3 p 2.826 N:O N1@A.G2648 O2@A.U2672
55 537 #4 p 2.965 O:N O2@A.C2649 N2@A.G2671
56 535 #5 p 2.836 N:N N3@A.C2649 N1@A.G2671
58 534 #6 p 2.769 N:O N4@A.C2649 O6@A.G2671
76 513 #7 p 2.806 N:N N3@A.U2650 N1@A.A2670
78 512 #8 p 3.129 O:N O4@A.U2650 N6@A.A2670
95 492 #9 p 2.703 O:N O2@A.C2651 N2@A.G2669
96 490 #10 p 2.853 N:N N3@A.C2651 N1@A.G2669
98 489 #11 p 2.987 N:O N4@A.C2651 O6@A.G2669
115 466 #12 p 2.817 O:N O2@A.C2652 N2@A.G2668
116 464 #13 p 2.907 N:N N3@A.C2652 N1@A.G2668
118 463 #14 p 2.897 N:O N4@A.C2652 O6@A.G2668
123 151 #15 o 2.622 O:O OP2@A.U2653 O2'@A.A2654
135 443 #16 p 2.898 O:N O2@A.U2653 N4@A.C2667
147 192 #17 x 3.054 O:O O4'@A.A2654 O4'@A.U2656
158 408 #18 p 2.960 N:O N6@A.A2654 OP2@A.C2666
173 188 #19 o 2.923 O:O O2'@A.G2655 OP2@A.U2656
173 378 #20 o 3.093 O:O O2'@A.G2655 O6@A.G2664
173 379 #21 o 3.343 O:N O2'@A.G2655 N1@A.G2664
181 386 #22 p 2.768 N:O N1@A.G2655 OP2@A.A2665
183 203 #23 p 2.754 N:O N2@A.G2655 O4@A.U2656
183 387 #24 p 2.887 N:O N2@A.G2655 O5'@A.A2665
188 379 #25 p 3.044 O:N OP2@A.U2656 N1@A.G2664
188 381 #26 p 2.944 O:N OP2@A.U2656 N2@A.G2664
200 401 #27 p 3.122 O:N O2@A.U2656 N6@A.A2665
201 398 #28 p 2.759 N:N N3@A.U2656 N7@A.A2665
220 381 #29 p 3.035 N:N N7@A.A2657 N2@A.G2664
223 371 #30 o 2.963 N:O N6@A.A2657 O2'@A.G2664
223 382 #31 p 3.039 N:N N6@A.A2657 N3@A.G2664
242 358 #32 p 2.821 O:N O2@A.C2658 N2@A.G2663
243 356 #33 p 2.890 N:N N3@A.C2658 N1@A.G2663
245 355 #34 p 2.887 N:O N4@A.C2658 O6@A.G2663
258 305 #35 o 2.604 O:N O2'@A.G2659 N7@A.A2661
258 308 #36 o 3.264 O:N O2'@A.G2659 N6@A.A2661
268 315 #37 p 2.973 N:O N2@A.G2659 OP2@A.A2662
268 327 #38 p 2.864 N:N N2@A.G2659 N7@A.A2662
371 390 #39 o 2.751 O:O O2'@A.G2664 O4'@A.A2665
550 566 #40 o 3.372 O:O O2'@A.U2672 O4'@A.G2673
In its default settings, DSSR detects 117 H-bonds for 1ehz (yeast phenylalanine tRNA), and 5,809 for 1jj2 (the H. marismortui large ribosomal subunit). Note that the program can identify H-bonds not only in RNA and DNA, but also in proteins, or their complexes. By default, however, DSSR only reports H-bonds within nucleic acids. As shown above, it is trivial to run DSSR with the --get-hbonds option to get all H-bonds in a given structure, and the plain text output is straightforward to work on.
While there exist dedicated tools for finding H-bonds, such as HBPLUS or HBexplore, DSSR may well be sufficient to fulfill most practical needs. If you notice any weird behaviors with this H-bond finding functionality, please let me know. I strive to address reported issues promptly, to the extent practical. At the very least, I should be able to explain why the program is working the way it does.

As of v1.0.3-2014mar09, DSSR has a decent user manual in PDF! Currently of 45 pages long, the DSSR manual contains everything a typical user needs to know to get started using the program effectively. The contents the manual are listed below.
Table of Contents
List of Figures
Introduction
Download and installation
Usages
Command-line help
Default run on PDB entry 1msy – detailed explanations
Summary section
List of base pairs
List of multiplets
List of helices
List of stems
List of lone canonical pairs
List of various loops
List of single-stranded fragments
Secondary structure in dot-bracket notation
List of backbone torsion angles and suite names
Default run on PDB entry 1ehz (tRNAPhe) – summary notes
Brief summary
Specific features
Default run on PDB entry 1jj2 – four auto-checked motifs
Kissing loops
A-minor (types I and II) motifs
Ribose zippers
Kink turns
The --more option
Extra parameters for base pairs
Extra parameters for helices/stems
The –-non-pair option
The –-u-turn option
The --po4 option
The –-long-idstr option
Frequently asked questions
How to cite DSSR?
Does DSSR work for DNA?
Does DSSR detect RNA tertiary interactions?
Revision history
Acknowledgements
References
With the User Manual available, I feel confident to claim that DSSR is now mature, stable, ready for real world applications. While only time would tell, I have no doubt that DSSR will become an essential tool in RNA structural bioinformatics.




![1msy [GUAA tetra loop] in 3d and 2d representations 1msy [GUAA tetra loop] in 3d and 2d representations](http://forum.x3dna.org/images/1msy-3d-2d.png)
