Once a nucleotide (nt) is identified, and matched to A (C, G, T, U) for the standard case or a (c, g, t, u) for a modified one, 3DNA/DSSR performs a least-squares fitting procedure to locate the base reference frame in three-dimensional space. The basic idea is very simple and widely applicable. The algorithm constitutes one of the key components of 3DNA/DSSR. As always, the details can be most effectively illustrated with a worked example. Using G1 in the yeast phenylalanine tRNA (PDB id: 1ehz) as an example, the atomic coordinates of its nine base-ring atoms are:
# G1, nine base-ring atoms for ls-fitting ATOM 14 N9 G A 1 51.628 45.992 53.798 1.00 93.67 N ATOM 15 C8 G A 1 51.064 46.007 52.547 1.00 92.60 C ATOM 16 N7 G A 1 51.379 44.966 51.831 1.00 91.19 N ATOM 17 C5 G A 1 52.197 44.218 52.658 1.00 91.47 C ATOM 18 C6 G A 1 52.848 42.992 52.425 1.00 90.68 C ATOM 20 N1 G A 1 53.588 42.588 53.534 1.00 90.71 N ATOM 21 C2 G A 1 53.685 43.282 54.716 1.00 91.21 C ATOM 23 N3 G A 1 53.077 44.429 54.946 1.00 91.92 N ATOM 24 C4 G A 1 52.356 44.836 53.879 1.00 92.62 C
The corresponding nine base-ring atoms of G in its standard base reference frame are listed below. See Table 1 of the report A Standard Reference Frame for the Description of Nucleic Acid Base-pair Geometry, and file Atomic_G.pdb distributed with 3DNA ($X3DNA/config/Atomic_G.pdb). In DSSR, the content has been integrated into the source code to make the program self-contained.
# G in standard base reference frame ATOM 2 N9 G A 1 -1.289 4.551 0.000 ATOM 3 C8 G A 1 0.023 4.962 0.000 ATOM 4 N7 G A 1 0.870 3.969 0.000 ATOM 5 C5 G A 1 0.071 2.833 0.000 ATOM 6 C6 G A 1 0.424 1.460 0.000 ATOM 8 N1 G A 1 -0.700 0.641 0.000 ATOM 9 C2 G A 1 -1.999 1.087 0.000 ATOM 11 N3 G A 1 -2.342 2.364 0.001 ATOM 12 C4 G A 1 -1.265 3.177 0.000
A least-squares fitting of the standard onto the experimental set of base-ring atoms defines the base reference frame (Fig. 1). The information is available via the following commands:
# find_pair -s 1ehz.pdb # in file 'ref_frames.dat'
... 1 G # A:...1_:[..G]G
53.7571 41.8678 52.9303 # origin
-0.2589 -0.2496 -0.9331 # x-axis
-0.5430 0.8365 -0.0731 # y-axis
0.7988 0.4878 -0.3521 # z-axis
# --------
# x3dna-dssr -i=1ehz.pdb --json | jq .nts[0].frame
{
rsmd: 0.008,
origin: [53.757, 41.868, 52.93],
x_axis: [-0.259, -0.25, -0.933],
y_axis: [-0.543, 0.837, -0.073],
z_axis: [0.799, 0.488, -0.352]
}
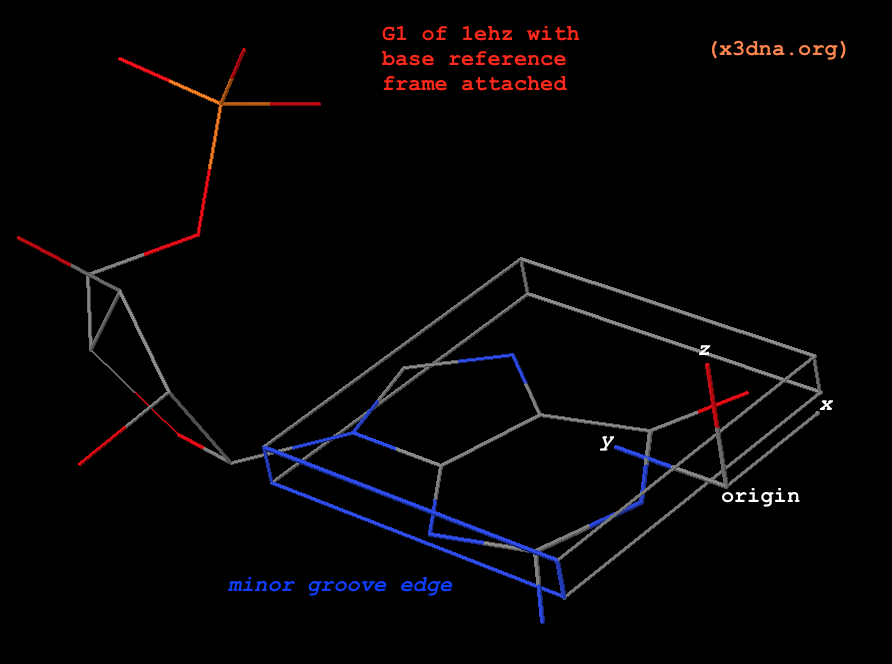
Fig. 1: G1 in tRNA 1ehz, with base reference frame attached
Please note the following subtle points:
- The standard base (
Atomic_G.pdb) is already set in its reference frame: the z-coordinates are virtually zeros, y-coordinates are positive, the atoms along the minor-groove edge have negative x-coordinates, as can be visualized clearly from the attached coordinate frame. In 3DNA, the five standard standard bases are in stored in filesAtomic_[ACGTU].pdb, and the corresponding modified ones are inAtomic_[acgtu].pdb. For simplicity,Atomic_A.pdbandAtomic_a.pdbare the same by default, as are the other four cases.
- The translation and rotation of the least-squares fitting process define the experimental base reference frame (for G1 in the above example), and its three axes are orthonormal by definition.
- By design, the base rings of
Atomic_A.pdbandAtomic_G.pdb match each other closely (see below), as are the pyrimidines bases. The least-square fitted root-mean-square deviation (rmsd) of the nine base-ring atoms between standard A and G is only 0.04 Å. Fitting the standard A (instead of G) onto G1 of 1ehz leads to a base reference frame that is essentially indistinguishable from the one above (see below). This feature shows that any ambiguity in assigning modified purines to A or G, or pyrimidines to C, T, or U causes no notable differences in 3DNA/DSSR results.
Comparison of base-ring atomic coordinates in standard G and A
Atomic_G.pdb Atomic_A.pdb
N9 G -1.289 4.551 0.000 | N9 A -1.291 4.498 0.000
C8 G 0.023 4.962 0.000 | C8 A 0.024 4.897 0.000
N7 G 0.870 3.969 0.000 | N7 A 0.877 3.902 0.000
C5 G 0.071 2.833 0.000 | C5 A 0.071 2.771 0.000
C6 G 0.424 1.460 0.000 | C6 A 0.369 1.398 0.000
N1 G -0.700 0.641 0.000 | N1 A -0.668 0.532 0.000
C2 G -1.999 1.087 0.000 | C2 A -1.912 1.023 0.000
N3 G -2.342 2.364 0.001 | N3 A -2.320 2.290 0.000
C4 G -1.265 3.177 0.000 | C4 A -1.267 3.124 0.000
Comparison of G1 (1ehz) base reference frame derived using standard G or A
Atomic_G.pdb | Atomic_A.pdb
53.7571 41.8678 52.9303 # origin | 53.7286 41.9276 52.9482 # origin
-0.2589 -0.2496 -0.9331 # x-axis | -0.2562 -0.2540 -0.9327 # x-axis
-0.5430 0.8365 -0.0731 # y-axis | -0.5444 0.8352 -0.0780 # y-axis
0.7988 0.4878 -0.3521 # z-axis | 0.7988 0.4878 -0.3522 # z-axis
