[Job] Staff Associate II (Computational Structural Biology) at Columbia University
The X3DNA-DSSR resource is at the forefront of structural bioinformatics, developing advanced tools for analyzing and modeling nucleic acid structures. We are seeking a highly motivated Staff Associate II to join our team and contribute to our next-generation analysis and visualization engine.
To see our resource in action, please visit wDSSR, our new web interface for dissecting and modeling 3D nucleic acid structures: https://web.x3dna-dssr.org/.
We are looking for a candidate with a strong scientific background in structural biology or bioinformatics and a desire to contribute to peer-reviewed publications through community-driven data analysis. We value individuals who are eager to learn, adapt to new technical challenges, and support the global research community.
For the full job description and to submit your application, please visit the official Columbia University posting:
https://apply.interfolio.com/183705
Announcing wDSSR: The Next-Generation Web Interface to X3DNA-DSSR
Dear 3DNA/DSSR Community,
We are thrilled to announce the official launch of wDSSR (https://web.x3dna-dssr.org/), the powerful new web interface to the X3DNA-DSSR analytical engine.
Developed by Drs. Shuxiang Li and Xiang-Jun Lu and supported by NIH grant R24GM153869, wDSSR represents a major leap forward from our highly popular 2019 Web 3DNA 2.0 framework. While Web 3DNA 2.0 has faithfully served the community for the analysis, visualization, and modeling of 3D nucleic acid structures, wDSSR was built from the ground up to take full advantage of modern web technologies and the latest DSSR backend capabilities.
A Modern, Streamlined Scientific Workflow
We have completely overhauled the user interface to provide a clean, intuitive, and task-driven experience. The core modeling and analysis tools are now seamlessly organized into a logical, single-word scientific workflow: Analyze, Rebuild, Model, Circularize, Mutate, Assemble, and Visualize.
Spotlight Feature: The "Assemble" Module
One of the most exciting upgrades is the newly renamed Assemble tab (formerly "Composite"). This advanced composite model builder allows you to effortlessly construct complex, higher-order models by linking any combination of nucleic acid duplexes or protein-DNA/RNA complexes. You can quickly connect up to six distinct target structures, ranging from simple linked A-DNA and B-DNA duplexes to large, protein-decorated structural assemblies.
Immediate Global Adoption
Although wDSSR has just launched, we are incredibly humbled to share that it is already seeing rapid worldwide adoption! According to recent network infrastructure data, the new interface is actively being used by researchers across North America, South America, Europe, and Asia. Within just a few days, we have recorded active sessions from prestigious institutions around the globe, including:
- The Weizmann Institute of Science in Israel
- Katholieke Universiteit Leuven in Belgium
- Queen's University in Canada
- Universidad Nacional Autonoma de Mexico (UNAM) in Mexico
- Emory University and the Wadsworth Centers Laboratories and Research in the United States
- Jawaharlal Nehru University and the China Education and Research Network in Asia
How to Cite
While a dedicated paper for wDSSR is currently in preparation, researchers should cite the server using its URL (https://web.x3dna-dssr.org/) alongside the 2019 Web 3DNA 2.0 paper and the foundational 2015 DSSR paper. Full details and funding acknowledgements can be found on our newly consolidated About page.
We invite you all to try out the new wDSSR platform! As always, your feedback is invaluable to us, and we encourage you to share your thoughts, questions, and structural models via the newly updated Questions & Feedback link in the wDSSR footer.
Happy modeling!
In the DSSR v1.2.7-2015jun09 release, I documented two additional command-line options (--prefix and --cleanup) that are related to the various auxiliary files. As a matter of fact, these two options (among quite a few others) have been there for a long time, but without being explicitly described. The point is not to hide but to simplify — one of the design goals of DSSR is simplicity. DSSR has already possessed numerous key functionality to be appreciated. Before DSSR is firmly established in the RNA bioinformatics field, I beleive too many nonessential “features” could be distracting. While writing and refining the DSSR code, I do feel that some ‘auxiliary’ features could be handy for experienced users (including myself). So along the way, I’ve added many ‘hidden’ options that are either experimental or potentially useful.
On one side, I sense it is acceptable for a scientific software to actually does more than it claims. On the other hand, I have always been quick in addressing users’ requests — as one example, check for the --select option recently introduced into DSSR in response to a user request, and the ‘hidden’ --dbn-break option for specifying the symbol to separate multiple chains or chain breaks in DSSR-derived dot-bracket notation.
Back to --prefix and --cleanup, the purposes of these two closely related options can be best illustrated using the yeast phenylalanine tRNA structure (1ehz) as an example. By default, running x3dna-dssr -i=1ehz.pdb will produce a total of 11 auxiliary files, with names prefixed with dssr-, as shown below:
List of 11 additional files
1 dssr-stems.pdb -- an ensemble of stems
2 dssr-helices.pdb -- an ensemble of helices (coaxial stacking)
3 dssr-pairs.pdb -- an ensemble of base pairs
4 dssr-multiplets.pdb -- an ensemble of multiplets
5 dssr-hairpins.pdb -- an ensemble of hairpin loops
6 dssr-junctions.pdb -- an ensemble of junctions (multi-branch)
7 dssr-2ndstrs.bpseq -- secondary structure in bpseq format
8 dssr-2ndstrs.ct -- secondary structure in connect table format
9 dssr-2ndstrs.dbn -- secondary structure in dot-bracket notation
10 dssr-torsions.txt -- backbone torsion angles and suite names
11 dssr-stacks.pdb -- an ensemble of stacks
With ‘fixed’ generic names by default, users can run DSSR in a directory repeatedly without creating too many files. This practice follows that used in the 3DNA suite of programs. However, my experience in supporting 3DNA over the years has shown that users (myself included) may want to explore further some of the files, e.g. ‘dssr-multiplets.pdb’ for displaying the base multiplets (four triplets here). One could easily use command-line (script) to change a generic name to a more appropriate one: e.g., mv dssr-multiplets.pdb 1ehz-multiplets.pdb for 1ehz. A better solution, however, is by introducing a customized prefix to the additional files, and that’s exactly where the --prefix option comes in. The option is specified like this: --prefix=text where text can be any string as appropriate. So running x3dna-dssr -i=1ehz.pdb --prefix=1ehz, for example, will lead to the following output:
List of 11 additional files
1 1ehz-stems.pdb -- an ensemble of stems
2 1ehz-helices.pdb -- an ensemble of helices (coaxial stacking)
3 1ehz-pairs.pdb -- an ensemble of base pairs
4 1ehz-multiplets.pdb -- an ensemble of multiplets
5 1ehz-hairpins.pdb -- an ensemble of hairpin loops
6 1ehz-junctions.pdb -- an ensemble of junctions (multi-branch)
7 1ehz-2ndstrs.bpseq -- secondary structure in bpseq format
8 1ehz-2ndstrs.ct -- secondary structure in connect table format
9 1ehz-2ndstrs.dbn -- secondary structure in dot-bracket notation
10 1ehz-torsions.txt -- backbone torsion angles and suite names
11 1ehz-stacks.pdb -- an ensemble of stacks
The --cleanup option, as its name implies, is to tidy up a directory by removing the auxiliary files generated by DSSR. The usage is very simple:
x3dna-dssr --cleanup
x3dna-dssr --cleanup --prefix=1ehz
The former gets rid of the default ‘fixed’ generic auxiliary files (dssr-pairs.pdb etc), whilst the latter deletes prefixed supporting files (1ehz-pairs.pdb etc).

Recently, I came across and have been surprised by the different assignment of HETATM vs. ATOM records for modified nucleotides in PDB vs. PDBx/mmCIF format. As always, the issue is best illustrated with a concrete example. Here is what I observed in the PDB entry 1ehz, the crystal structure of yeast phenylalanine tRNA at 1.93 Å resolution.
DSSR identifies 14 modified nucleotides (of 11 types) in 1ehz as shown below:
List of 11 types of 14 modified nucleotides
nt count list
1 1MA-a 1 A.1MA58
2 2MG-g 1 A.2MG10
3 5MC-c 2 A.5MC40,A.5MC49
4 5MU-t 1 A.5MU54
5 7MG-g 1 A.7MG46
6 H2U-u 2 A.H2U16,A.H2U17
7 M2G-g 1 A.M2G26
8 OMC-c 1 A.OMC32
9 OMG-g 1 A.OMG34
10 PSU-P 2 A.PSU39,A.PSU55
11 YYG-g 1 A.YYG37
In file 1ehz.pdb downloaded from RCSB PDB, all the 14 modified nucleotides are assigned as HETATM whereas in 1ehz.cif the corresponding records are ATOM. Here is the excerpt for 1MA58 in PDB format:
HETATM 1252 P 1MA A 58 73.770 67.765 34.057 1.00 30.65 P
HETATM 1253 OP1 1MA A 58 72.638 67.886 33.105 1.00 32.84 O
HETATM 1254 OP2 1MA A 58 73.621 68.229 35.450 1.00 29.49 O
HETATM 1255 O5' 1MA A 58 74.315 66.273 34.254 1.00 28.81 O
HETATM 1256 C5' 1MA A 58 74.592 65.439 33.080 1.00 29.42 C
HETATM 1257 C4' 1MA A 58 74.279 63.972 33.383 1.00 33.42 C
HETATM 1258 O4' 1MA A 58 74.880 63.685 34.667 1.00 32.36 O
HETATM 1259 C3' 1MA A 58 72.789 63.573 33.509 1.00 35.13 C
HETATM 1260 O3' 1MA A 58 72.625 62.168 33.250 1.00 36.80 O
HETATM 1261 C2' 1MA A 58 72.560 63.667 35.012 1.00 34.80 C
HETATM 1262 O2' 1MA A 58 71.525 62.828 35.506 1.00 36.27 O
HETATM 1263 C1' 1MA A 58 73.908 63.150 35.551 1.00 33.62 C
HETATM 1264 N9 1MA A 58 74.284 63.494 36.930 1.00 30.36 N
HETATM 1265 C8 1MA A 58 73.887 64.574 37.688 1.00 34.55 C
HETATM 1266 N7 1MA A 58 74.415 64.610 38.899 1.00 33.32 N
HETATM 1267 C5 1MA A 58 75.204 63.469 38.953 1.00 33.37 C
HETATM 1268 C6 1MA A 58 76.031 62.941 39.948 1.00 33.58 C
HETATM 1269 N6 1MA A 58 76.184 63.488 41.134 1.00 41.19 N
HETATM 1270 N1 1MA A 58 76.708 61.803 39.669 1.00 34.48 N
HETATM 1271 CM1 1MA A 58 77.649 61.222 40.626 1.00 31.43 C
HETATM 1272 C2 1MA A 58 76.527 61.216 38.479 1.00 28.43 C
HETATM 1273 N3 1MA A 58 75.793 61.624 37.453 1.00 31.67 N
HETATM 1274 C4 1MA A 58 75.142 62.771 37.747 1.00 33.02 C
The corresponding section in PDBx/mmCIF format is:
ATOM 1252 P P . 1MA A 1 58 ? 73.770 67.765 34.057 1.00 30.65 ? ? ? ? ? ? 58 1MA A P 1
ATOM 1253 O OP1 . 1MA A 1 58 ? 72.638 67.886 33.105 1.00 32.84 ? ? ? ? ? ? 58 1MA A OP1 1
ATOM 1254 O OP2 . 1MA A 1 58 ? 73.621 68.229 35.450 1.00 29.49 ? ? ? ? ? ? 58 1MA A OP2 1
ATOM 1255 O "O5'" . 1MA A 1 58 ? 74.315 66.273 34.254 1.00 28.81 ? ? ? ? ? ? 58 1MA A "O5'" 1
ATOM 1256 C "C5'" . 1MA A 1 58 ? 74.592 65.439 33.080 1.00 29.42 ? ? ? ? ? ? 58 1MA A "C5'" 1
ATOM 1257 C "C4'" . 1MA A 1 58 ? 74.279 63.972 33.383 1.00 33.42 ? ? ? ? ? ? 58 1MA A "C4'" 1
ATOM 1258 O "O4'" . 1MA A 1 58 ? 74.880 63.685 34.667 1.00 32.36 ? ? ? ? ? ? 58 1MA A "O4'" 1
ATOM 1259 C "C3'" . 1MA A 1 58 ? 72.789 63.573 33.509 1.00 35.13 ? ? ? ? ? ? 58 1MA A "C3'" 1
ATOM 1260 O "O3'" . 1MA A 1 58 ? 72.625 62.168 33.250 1.00 36.80 ? ? ? ? ? ? 58 1MA A "O3'" 1
ATOM 1261 C "C2'" . 1MA A 1 58 ? 72.560 63.667 35.012 1.00 34.80 ? ? ? ? ? ? 58 1MA A "C2'" 1
ATOM 1262 O "O2'" . 1MA A 1 58 ? 71.525 62.828 35.506 1.00 36.27 ? ? ? ? ? ? 58 1MA A "O2'" 1
ATOM 1263 C "C1'" . 1MA A 1 58 ? 73.908 63.150 35.551 1.00 33.62 ? ? ? ? ? ? 58 1MA A "C1'" 1
ATOM 1264 N N9 . 1MA A 1 58 ? 74.284 63.494 36.930 1.00 30.36 ? ? ? ? ? ? 58 1MA A N9 1
ATOM 1265 C C8 . 1MA A 1 58 ? 73.887 64.574 37.688 1.00 34.55 ? ? ? ? ? ? 58 1MA A C8 1
ATOM 1266 N N7 . 1MA A 1 58 ? 74.415 64.610 38.899 1.00 33.32 ? ? ? ? ? ? 58 1MA A N7 1
ATOM 1267 C C5 . 1MA A 1 58 ? 75.204 63.469 38.953 1.00 33.37 ? ? ? ? ? ? 58 1MA A C5 1
ATOM 1268 C C6 . 1MA A 1 58 ? 76.031 62.941 39.948 1.00 33.58 ? ? ? ? ? ? 58 1MA A C6 1
ATOM 1269 N N6 . 1MA A 1 58 ? 76.184 63.488 41.134 1.00 41.19 ? ? ? ? ? ? 58 1MA A N6 1
ATOM 1270 N N1 . 1MA A 1 58 ? 76.708 61.803 39.669 1.00 34.48 ? ? ? ? ? ? 58 1MA A N1 1
ATOM 1271 C CM1 . 1MA A 1 58 ? 77.649 61.222 40.626 1.00 31.43 ? ? ? ? ? ? 58 1MA A CM1 1
ATOM 1272 C C2 . 1MA A 1 58 ? 76.527 61.216 38.479 1.00 28.43 ? ? ? ? ? ? 58 1MA A C2 1
ATOM 1273 N N3 . 1MA A 1 58 ? 75.793 61.624 37.453 1.00 31.67 ? ? ? ? ? ? 58 1MA A N3 1
ATOM 1274 C C4 . 1MA A 1 58 ? 75.142 62.771 37.747 1.00 33.02 ? ? ? ? ? ? 58 1MA A C4 1
While I have not tested exhaustively, it seems true that PDBx/mmCIF has adopted a different definition of what constitutes a HETATM residue. It is worth noting that results from 3DNA and DSSR/SNAP are not effected by the conflicting assignments.

Nowadays, “big data” and “big science” are hot topics. They all sound good and certainly come about for a reason. Yet, to transform data to information to knowledge to understanding to wisdom, sophisticated software tools are required. The programs can be big and complicated, or small and self-contained, fitting different purposes. As long as they can get the claimed job done in a robust fashion, size should not be a concern.
Over the years, however, I have seen a trend of bloated software with many (fragile) dependencies in bioinformatics. Some tools are so picky and hard to use/maintain that instead of serving, they become sort of a master. As a more representative example, I recently tried to install an open-source software associated with a paper published just a few years ago in a leading journal. The software has only a few dependencies, yet some of them have already become obsolete. I spent hours each time, on Mac OS X and two versions of Ubuntu Linux, but failed to get it running properly (always abort with error messages). The download page hosting the software has been inactive since around the publication of the paper. Presumably, the PhD student or postdoc who wrote the code had left the lab, and with a paper published, all is done!
As an active practitioner of bioinformatics for well over a decade, I can confidently claim to be well above average in familiarity with Linux/Mac OS X and associated shell programming and make etc tools, and various common scripting and compiled programming languages. Yet, once in a while, I get frustrated when I try to download and install a software tool attached to a paper I am interested in. As I see it, the vast majority of software programs from research labs are publication-oriented — as long a paper is published, it is finished.
From my experience, I always see software as engineering. It needs careful design and great attention to meticulous details. A sophisticated piece of scientific software is a combination of science and engineering. Expertise in domain knowledge is a must, and refined skills in computer programming is indispensable. The DSSR program I created and continuously refined over the past three years represents what a scientific software should be in my believe.
Among other unique features, DSSR is tiny (< 1mb), self-contained (without run-time dependencies) and runs on Windows, Mac OS X, and Linux. Getting DSSR up and running should take only minutes by any one with basic familiarity of common computer systems. I have no doubt that the beauty of being small as represented by DSSR will be gradually appreciated by the community.

Over the past few weeks, I’ve had the pleasure to talk to Thomas Holder, the PyMOL Principal Developer at Schrödinger, on possible integration of DSSR into PyMOL. On Tuesday April 21, 2015, I wrote to Thomas:
Last year, I had the please to collaborate with Dr. Robert Hanson to integrate DSSR into Jmol, see
http://chemapps.stolaf.edu/jmol/jsmol/dssr.htm. I am wondering if you have any interest in connecting DSSR to PyMOL. This will not only benefit both parties, but also bring elaborate analyses of RNA structures to the general audience. As you may be aware, RNA is becoming increasing important, yet the field of RNA structural bioinformatics is lagging (far) behind that of proteins.
After a few meet-ups, we all agree that the DSSR-PyMOL integration project would be meaningful/significant for RNA structural bioinformatics. Moreover, the community not only can benefit from the end result, but also should be able to make direct contributions through the process. On Friday May 08, 2015, Thomas sent out the following open invitation, titled Someone interested in writing a DSSR plugin for PyMOL?, to the PyMOL mailing list:
Is anyone interested in writing a DSSR plugin for PyMOL? DSSR is an integrated software tool for Dissecting the Spatial Structure of RNA (http://x3dna.bio.columbia.edu/docs/dssr-manual.pdf). Among other things, DSSR defines the secondary structure of RNA from 3D atomic coordinates in a way similar to DSSP does for proteins. Most of its output could be translated 1:1 into PyMOL selections, making it available for coloring and other selection based features. A PyMOL plugin could act as a wrapper which runs DSSR for an object or atom selection. Xiang-Jun Lu, the author of DSSR, is also working on base pair visualization (see http://x3dna.org/articles/seeing-is-understanding-as-well-as-believing), similar to (but more advanced) what’s already available from 3DNA (http://pymolwiki.org/index.php/3DNA).
Xiang-Jun would be happy to collaborate with someone who has experience with Python and the PyMOL API for writing an extension or plugin. Please contact me if this sounds appealing to you.
Get DSSR from http://x3dna.org/
See it hooked up with JSmol: http://chemapps.stolaf.edu/jmol/jsmol/dssr.htm
If you are self-motivated, care about software quality, have expertise in writing PyMOL plugin, and feel the pain in RNA structural analysis/visualization with currently available tools, now it is the time to make a difference. The DSSR/PyMOL project would ideally be composed of a team of dedicated practitioners with complementary skills. We will communicate mostly via email or online forum, in a presumably open and highly interactive way. By working on the project, you will be able to sharpen your skills and make new friends. The end product would not only make RNA structural bioinformatics easier for yourself but also benefit the community at large.

From the very beginning, 3DNA contains two key programs, analyze and rebuild, for the analysis and rebuilding of nucleic acid 3D structures. The two names are short and to the point, but with one caveat. They are common verbs that can be easily picked up by other software packages. When 3DNA and such packages are installed on the same machine, naming clashes happen. If the 3DNA bin/ directory is searched afterwards, the analyze or rebuild command may have nothing to do with nucleic acid structures at all. Naturally, this naming ambiguity can lead to confusions and frustrations.
I’ve been aware of the rebuild program name conflict for a long time. Recently, I was surprised by another analyze program on my Mac OS X Yosemite. As shown from the following output, the analyze program seems to be installed via Mac port, and it is about analyzing words in a dictionary file.
~ [540] which analyze
/opt/local/bin/analyze
~ [541] analyze -h
correct syntax is:
analyze affix_file dictionary_file file_of_words_to_check
use two words per line for morphological generation
The ambiguous names are exactly the reason that I use x3dna-dssr and x3dna-snap for the two new programs I’ve been working over the past few years. As for the analyze and rebuild programs in 3DNA v2.x, I’d rather leave them as is. 3DNA is now in wide use in other structural bioinformatic pipelines to allow for easy name changes without causing compatibility issues. On a positive side, once you know the problem, fixing it is straightforward. This post is to raise the awareness of the 3DNA user community about such naming conflicts.

Canonical bases (A, C, G, T and U) in nucleic acid structures have standard atom names, shown below using the Watson-Crick A–T and G–C pairs. Ring atoms of adenine, for example, are named (N1, C2, N3, C4, C5, C6, N7, C8, N9) respectively.
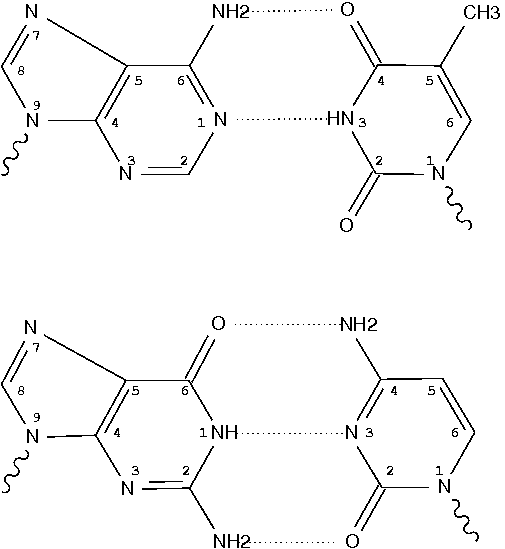
Four characters are reserved for atom names in the PDB format. The convention, as seen in files downloaded from the RCSB PDB, is to put the two-character base name in the middle, as in .N1.. Note that here each dot (.) is used for a space character to make it stand out.
Long time ago, I became aware a PDB format variant where the base name is left-aligned, as in N1... This case has ever since been properly handled by 3DNA (including DSSR and SNAP). While checking submitted entries to web-DSSR, I recently noticed yet another PDB format variation in labeling base names with the format of ..N1 (i.e., right-aligned). Without taking this special variant of PDB format into consideration, 3DNA/DSSR reported that “no nucleotides found!” Once the issue is known, however, fixing it is straightforward. As of May 4, 2015, 3DNA v2.2, DSSR and SNAP can all handle this special PDB variant correctly.
Over the years, I have come across many PDB variants claimed to compliant with the loosely defined format. If you find 3DNA or DSSR is not working as expected, it is likely the coordinate file in the self-claimed ‘PDB format’ is at fault. Wherever practical, I’ve tried to incorporate as many non-standard variants as possible.

The NDB (Nucleic Acid Database) is a valuable resource dedicated to “information about experimentally-determined nucleic acids and complex assemblies.” Over the years, however, I’ve gradually switched from NDB to PDB (Protein Data Bank) for my research on nucleic acid structures. NDB is derived from PDB and presumably should contain all nucleic acid structures available in the PDB. However, at the time of this writing (on April 9, 2015), the NDB says: “As of 8-Apr-2015 number of released structures: 7430” and the PDB states “7611 Nucleic Acid Containing Structures”. So PDB has 7611-7430=181 more entries of nucleic acid structures than the NDB, possibly due to a lag in NDB’s processing of newly released PDB structures. Another issue is the inconsistency of the NDB identifier: early entries have e.g. bdl084 for B-DNA (355d in PDB), but now NDB seems to use the same id as the PDB (e.g., 4p5j).
The RCSB PDB maintains a weekly-updated, summary file named pdb_entry_type.txt in pure text format (check here for a list of useful summary files), containing “List of all PDB entries, identification of each as a protein, nucleic acid, or protein-nucleic acid complex and whether the structure was determined by diffraction or NMR.” An excerpt of the file is shown below:
108m prot diffraction
109d nuc diffraction
109l prot diffraction
109m prot diffraction
10gs prot diffraction
10mh prot-nuc diffraction
110d nuc diffraction
110l prot diffraction
.................................
102m prot diffraction
103d nuc NMR
Specifically, a nucleic acid structure contains the (sub)string nuc in the second field, where prot-nuc means a protein-RNA/DNA complex. This text file is trivial to parse, and the atomic coordinates files (in PDB or PDBx/mmCIF format) for all nucleic acid structures can be automatically downloaded from the RCSB PDB using a script.
It is worth noting that DSSR is checked against all nucleic acid structures in the PDB at the time of each release to ensure that it does not crash. I update my local copy of nucleic acid structures each week, and run DSSR on the new entries. This process not only provides me an opportunity to keep pace with new developments in the field but also allows me to keep refining DSSR as needs arise.

Pseudouridine (5-ribosyluracil, PSU) is the most abundant modified nucleotide in RNA. It is unique in that it has a C-glycosidic bond (C-C1′) instead of the N-glycosidic bond (N-C’) common to all other nucleotides, canonical or modified. In 3DNA, the one-letter code for PSU is assigned to the upper case ‘P’, reserving the lower case ‘p’ for its modified variants. Distinguishing PSU from standard U (or T) is important for deriving sensible base-pair parameters and the χ torsion angle.
 |
 |
|
| PSU |
3TD |
Recently, I came across 3TD (see figure above) in PDB entry 5afi. 3TD is a modified variant of PSU, with a methyl group attached to N3. In 3DNA v2.1 v2.1-2015mar11, 3TD is abbreviated to ‘p’ to signify its connection to PSU.
In the list of recognized nucleotides (‘baselist.dat’) distributed with 3DNA, there are two other residues mapped to ‘p’: FHU and P2U (see figure below). As is often the case, it is the chemical structure, not the 3-letter PDB ligand identifier (or even full chemical name), that shows clearly to what 3DNA 1-letter abbreviation a residue matches.
 |
 |
|
| FHU |
P2U |

A single-stranded RNA molecule can fold back onto itself to form various loops delineated by double helical stems, as shown in the figure below [taken from the Nearest Neighbor Database website from the Turner group].
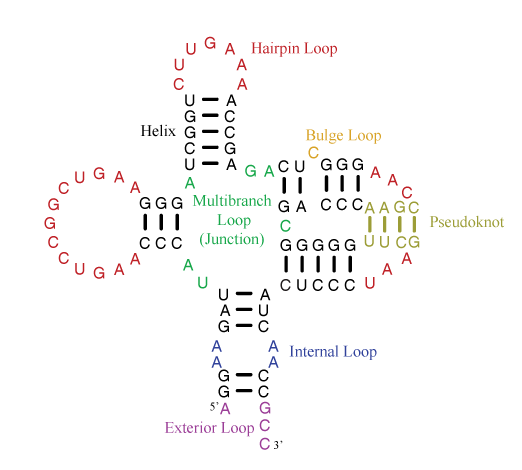
Of special note is the exterior loop (at the bottom) which includes the 5′ and 3′ ends of the sequence. The Mfold User Manual defines the exterior loop as such:
The collection of bases and base pairs not accessible from any base pair is called the exterior (or external) loop … . It is worth noting that if we imagine adding a 0th and an (n + 1)st base to the RNA, and a base pair 0.(n+1), then the exterior loop becomes the loop closed by this imaginary base pair. … The exterior loop exists only in linear RNA.
While each of the other loops (hairpin, bulge, internal or junction) forms a closed ‘circle’ with two neighboring bases connected by either a canonical pair or backbone covalent bond, the ‘exterior loop’ has only an imaginary pair to close the 5′ and 3′ ends of the sequence. Moreover, the two ends of an RNA molecule are not necessarily close in three-dimenional space, as may be implied in the above secondary structure diagram. For example, in the H-type pseudoknotted structure 1ymo from human telomerase RNA, the 5′ and 3′ ends are on the opposite sides.
DSSR does not has the concept of an ‘exterior loop’ due to its lack of a closing pair to form a ‘circle’. Instead, each of the 5′ and 3′ dangling ends is taken as a ‘non-loop single-stranded segment’, if applicable. For the crystal structure of yeast phenylalanine tRNA (1ehz, see the figure at the bottom), the relevant portion of DSSR output is as below. Note that since the 5′ end is paired, only the single-stranded region at the 3′ end is listed. Presumably, the ‘exterior loop’ in this case would also include the G1—C72 pair, with the imaginary closing link connecting G1 and A76.
List of 1 non-loop single-stranded segment
1 nts=4 ACCA A.A73,A.C74,A.C75,A.A76









