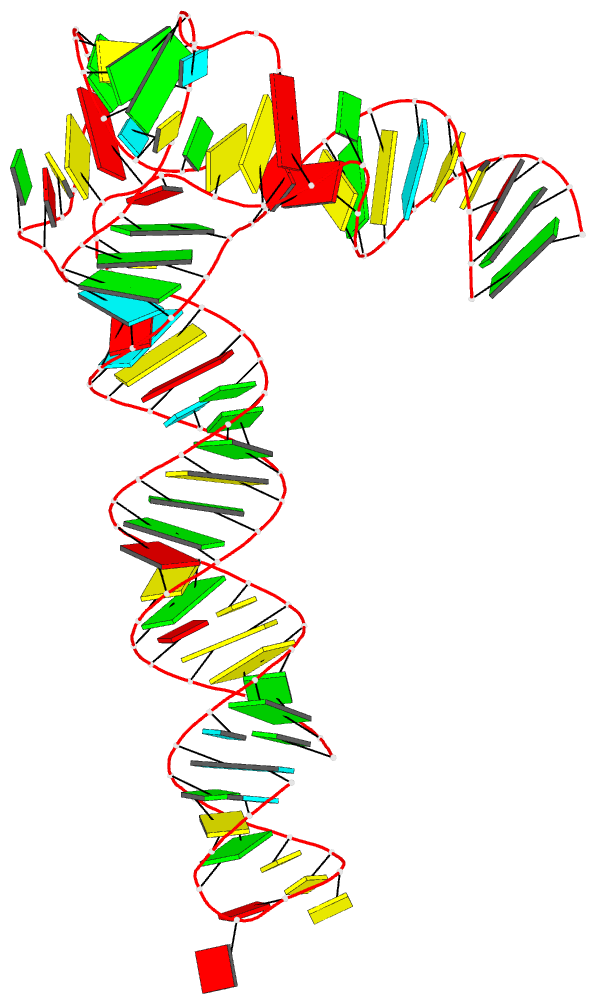
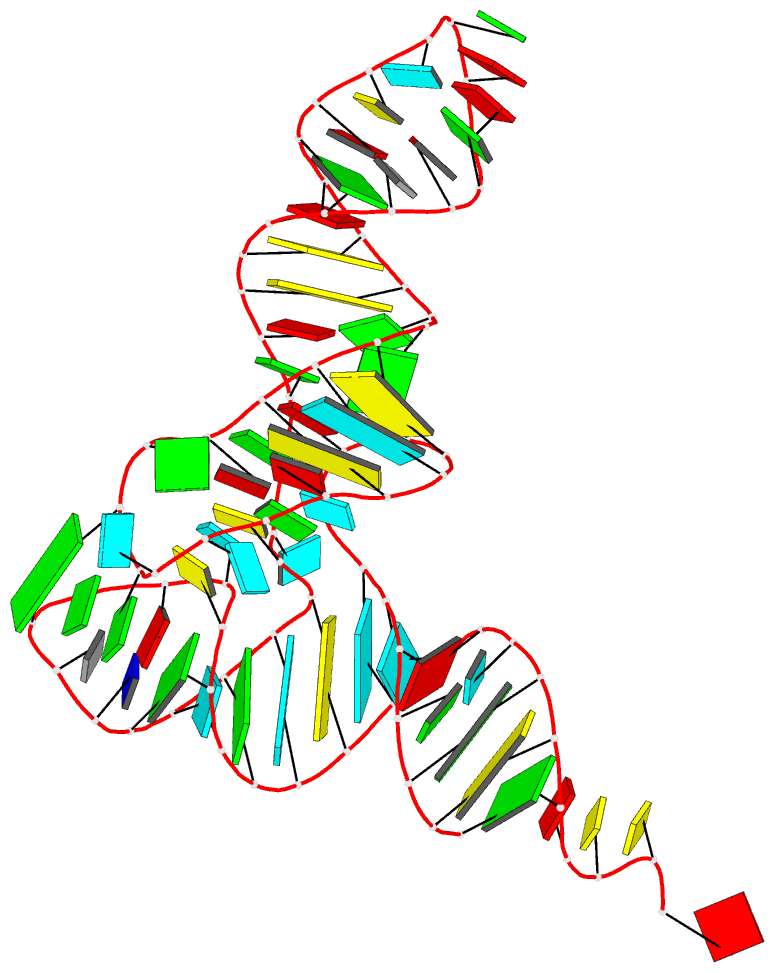
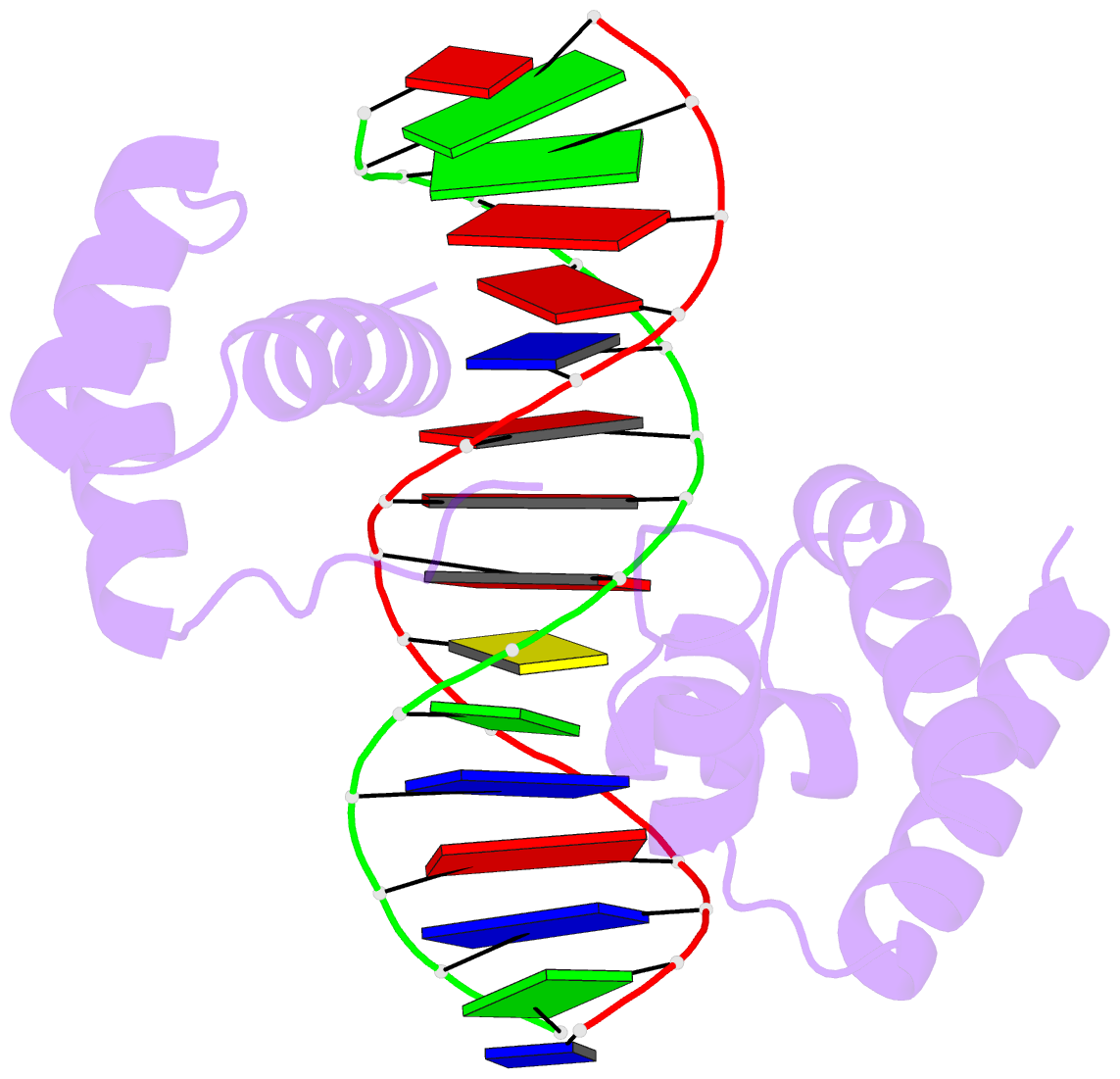
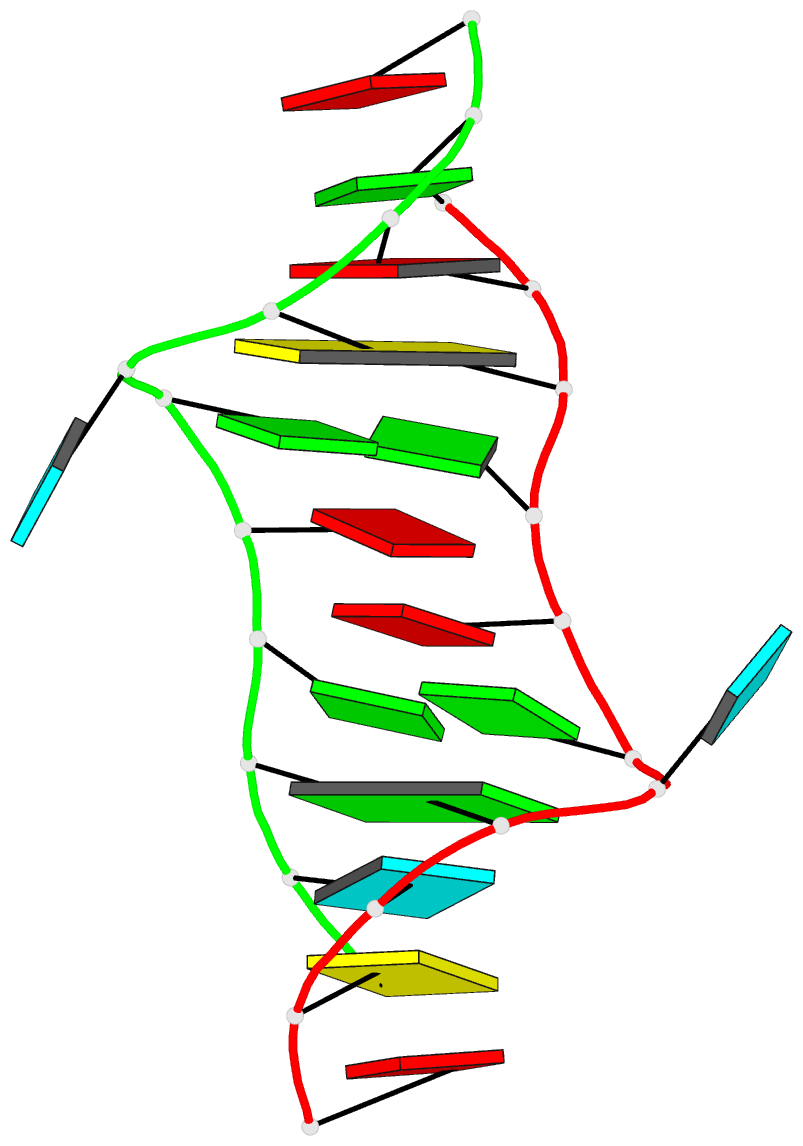
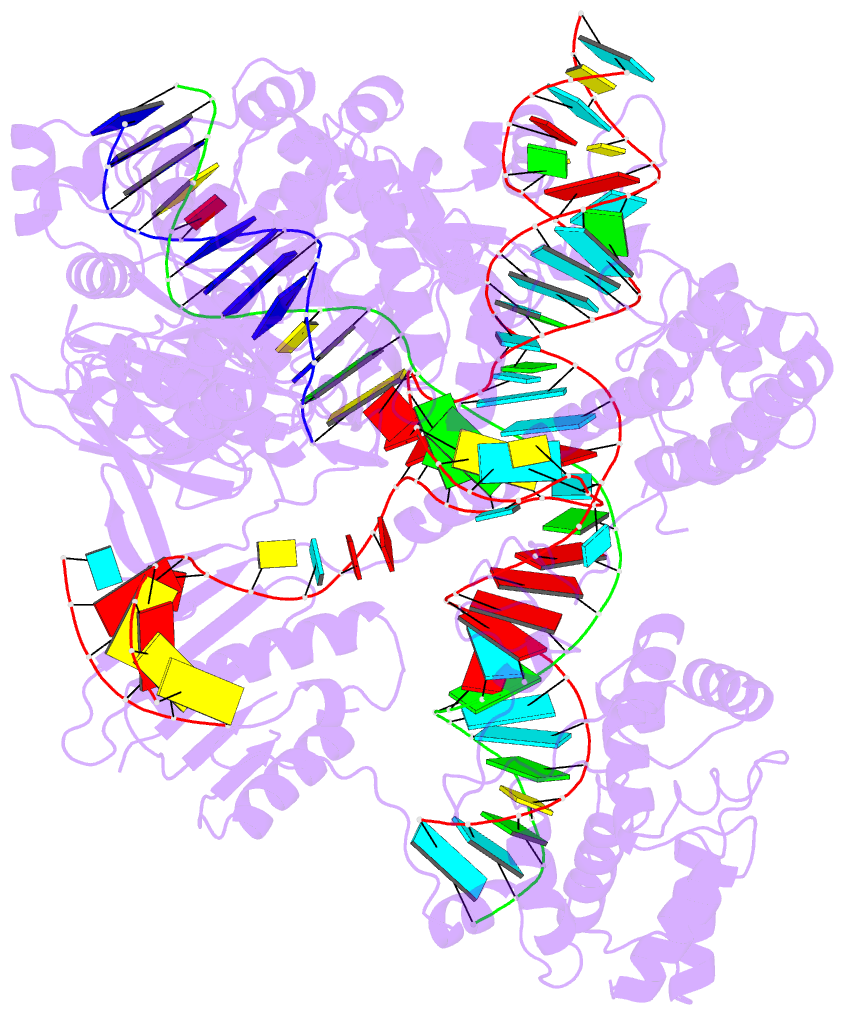
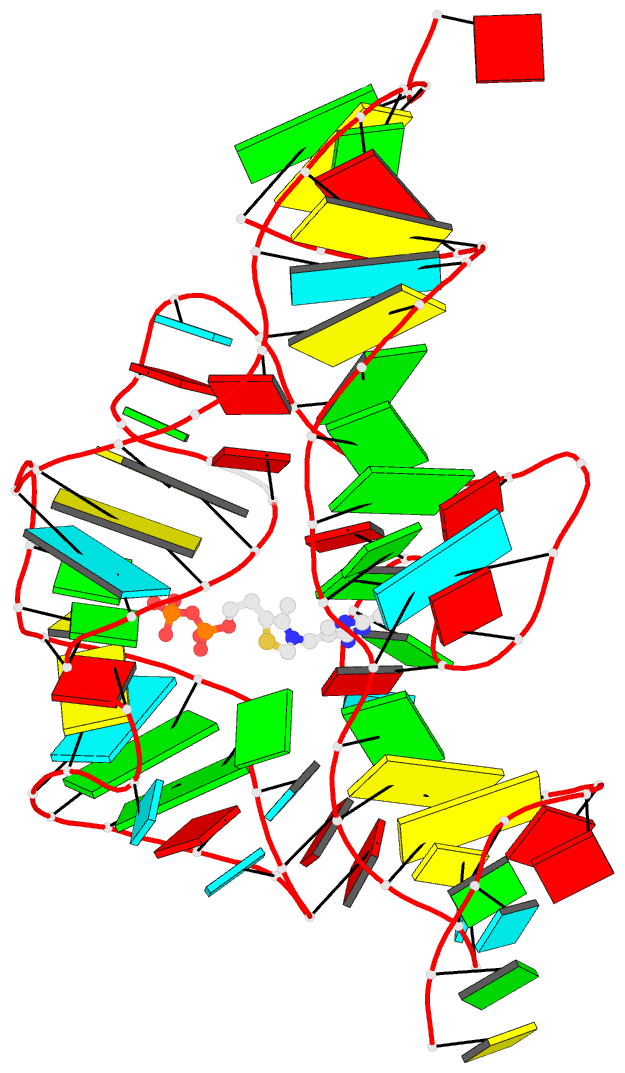
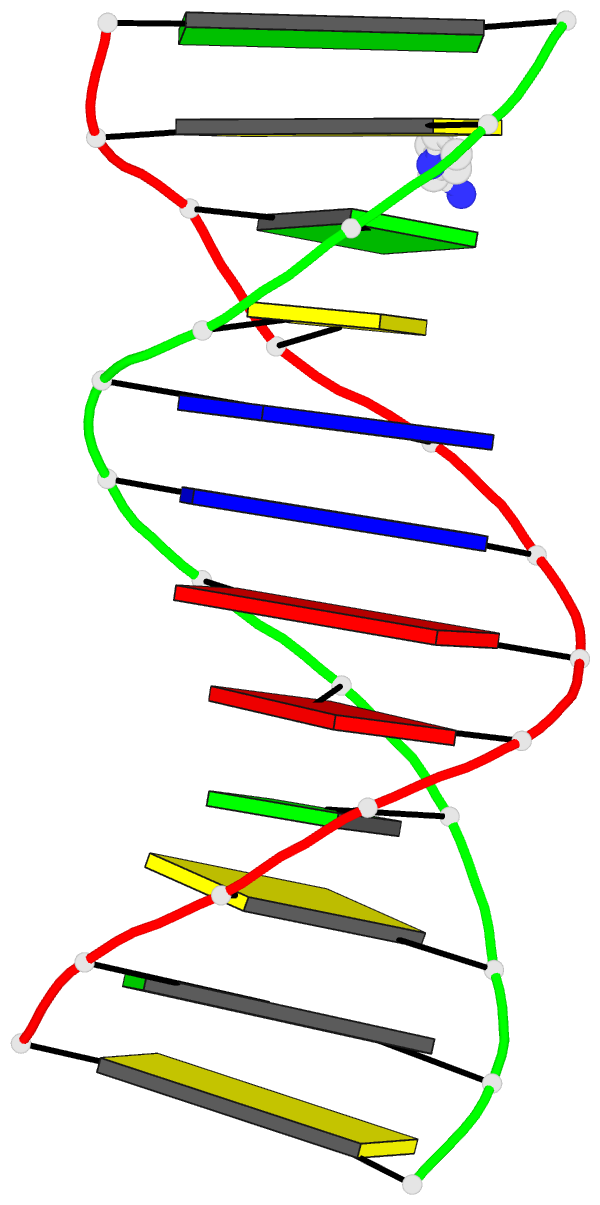
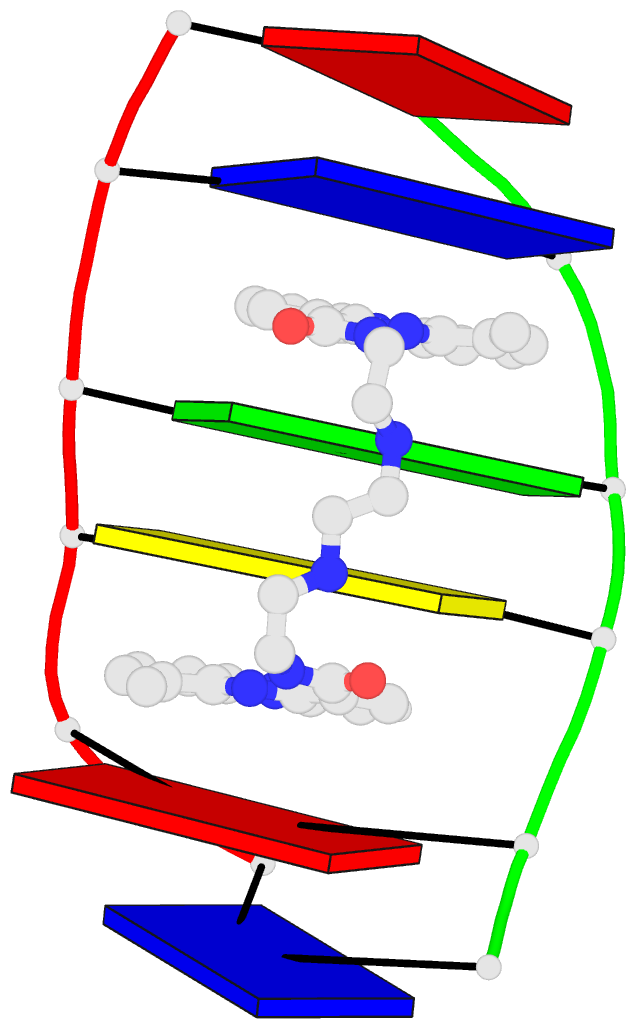
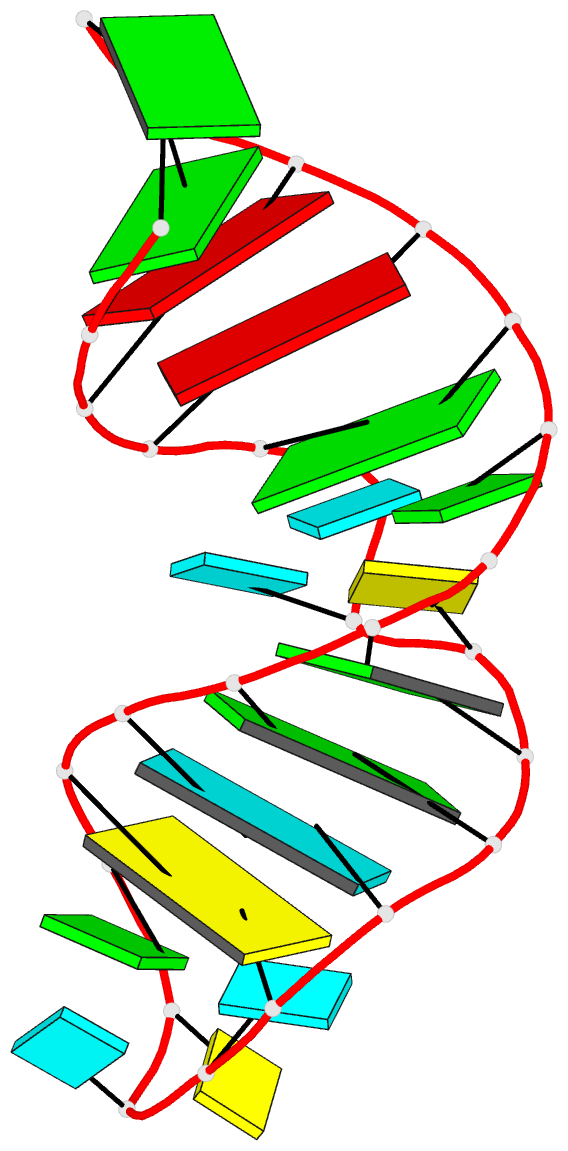
The paper, titled DSSR-enabled innovative schematics of 3D nucleic acid structures with PyMOL, has just been published in Nucleic Acids Research (online on May 22, 2020). Here is the abstract:
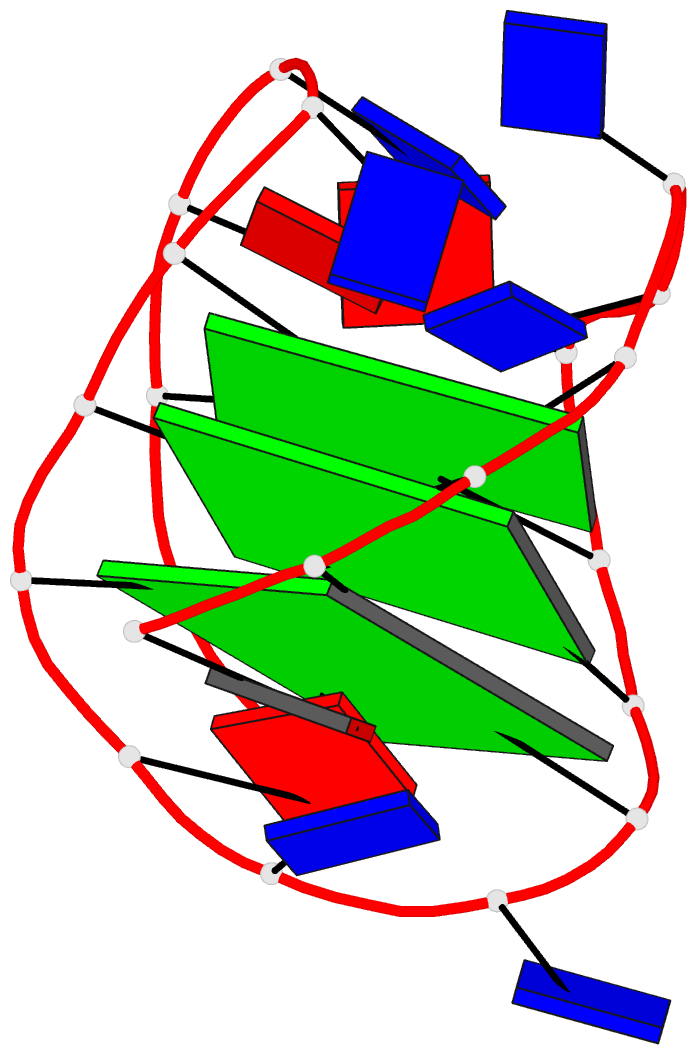
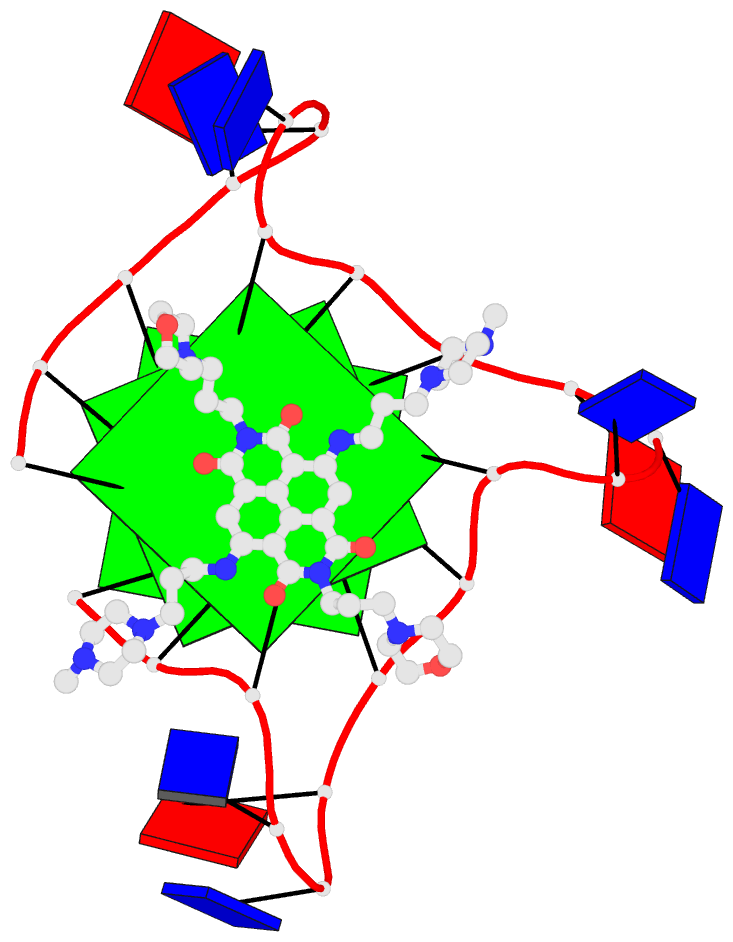
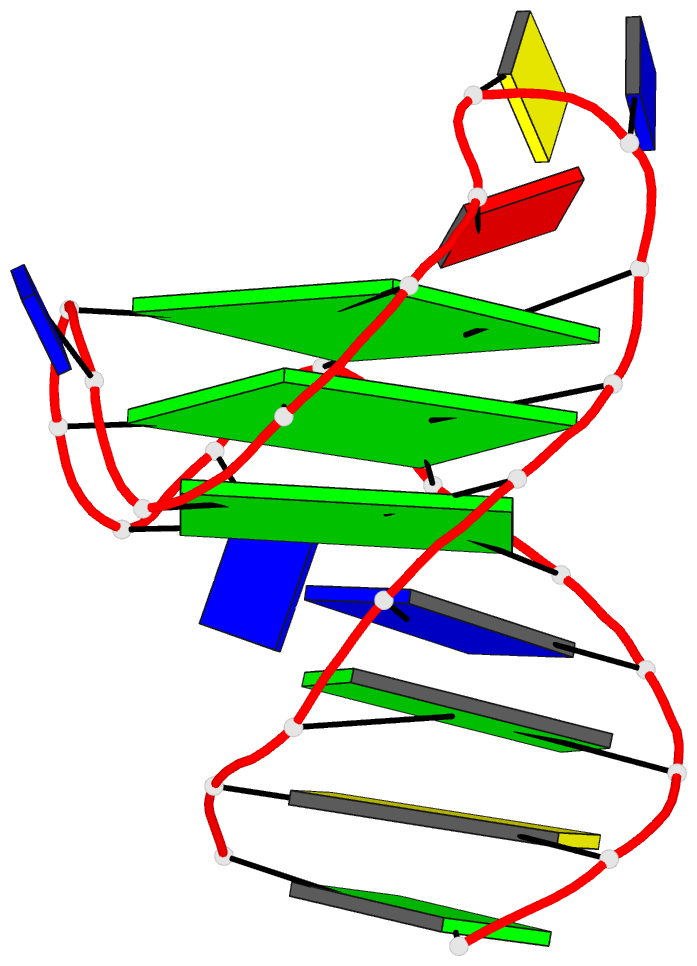
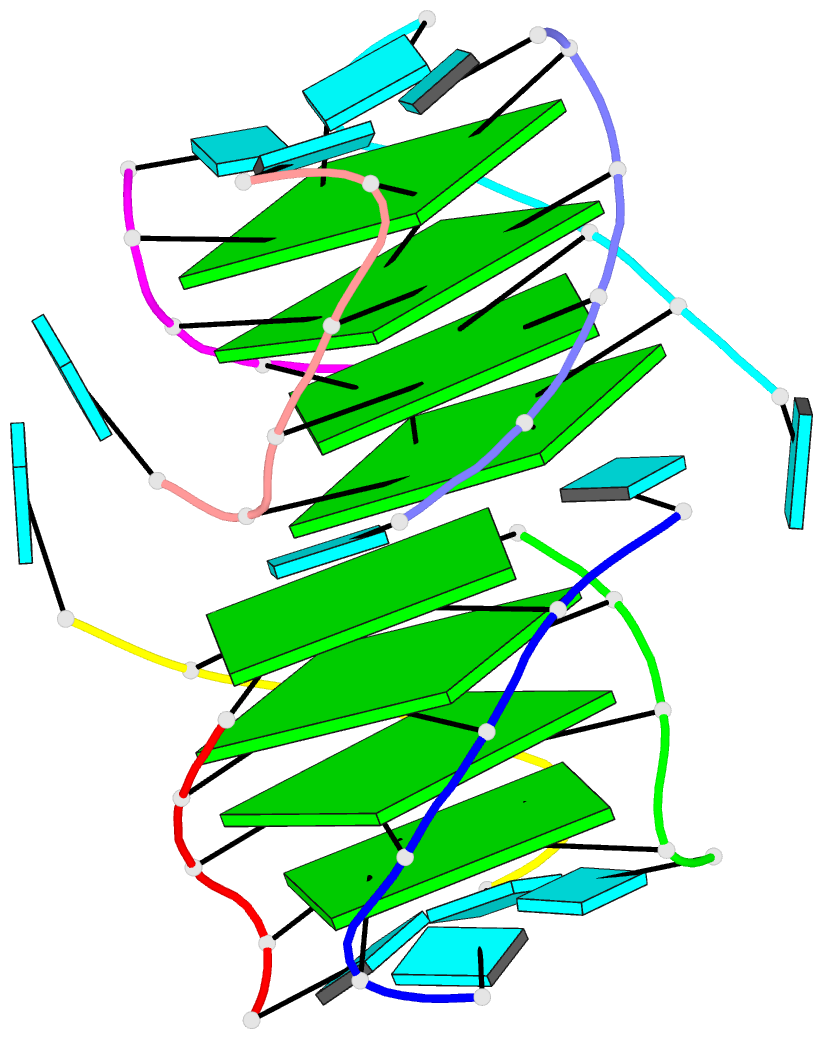
Sophisticated analysis and simplified visualization are crucial for understanding complicated structures of biomacromolecules. DSSR (Dissecting the Spatial Structure of RNA) is an integrated computational tool that has streamlined the analysis and annotation of 3D nucleic acid structures. The program creates schematic block representations in diverse styles that can be seamlessly integrated into PyMOL and complement its other popular visualization options. In addition to portraying individual base blocks, DSSR can draw Watson-Crick pairs as long blocks and highlight the minor-groove edges. Notably, DSSR can dramatically simplify the depiction of G-quadruplexes by automatically detecting G-tetrads and treating them as large square blocks. The DSSR-enabled innovative schematics with PyMOL are aesthetically pleasing and highly informative: the base identity, pairing geometry, stacking interactions, double-helical stems, and G-quadruplexes are immediately obvious. These features can be accessed via four interfaces: the command-line interface, the DSSR plugin for PyMOL, the web application, and the web application programming interface. The supplemental PDF serves as a practical guide, with complete and reproducible examples. Thus, even beginners or occasional users can get started quickly, especially via the web application at http://skmatic.x3dna.org.
A brief history on DNA/RNA schematics as implemented in SCHNAaP/SCHNArP, 3DNA, and now in DSSR:
The idea of representing bases and WC-pairs as rectangular blocks came from the pioneering work of Calladine et al. (27,28) The block schematics were first implemented in the pair of SCHNAaP/SCHNArP programs (29,30) for rigorous analysis and reversible rebuilding of double-helical nucleic acid structures. The algorithms that underpinned SCHNAaP/SCHNArP laid the foundation of ‘analyze’ and ‘rebuild’, two core components of the 3DNA suite of programs (31–33). 3DNA also takes advantage of the standard base reference frame (34), and comprises quite a few other related programs. One of them is ‘blocview’, a script which calls several 3DNA utility programs to generate individual base blocks and set the view, MolScript (35) to produce backbone ribbons, and Raster3D (36) to render the composite image. The 3DNA ‘blocview’ schematics catch characteristic attributes of nucleic acid structures. They have gradually become popular and been adopted into the RCSB PDB (1) and the NDB (37), and then propagated into other bioinformatics resources (e.g., the ‘RNA Structure Atlas’ website hosted by the Leontis-Zirbel RNA group).
DSSR supersedes ‘blocview’ by eliminating all the internal and external dependencies of the 3DNA utility program. DSSR produces block representations, not only of individual bases but also WC-pairs and G-tetrads, that can be fed directly into PyMOL. The DSSR-PyMOL integration is easier to use, has more features, and produces better schematics than the original 3DNA-blocview approach.
Indeed, the base block schematics have continuously evolved for over two decades, as appreciated in the acknowledgements:
I would like to thank Christopher A. Hunter, Christopher R. Calladine, Helen M. Berman, Catherine L. Lawson, Zukang Feng, Wilma K. Olson and Harmen J. Bussemaker for their helpful input on the block schematic during its continuous evolution for over two decades. I appreciate Thomas Holder (PyMOL Principal Developer, Schrödinger, Inc.) for writing the DSSR plugin for PyMOL, and for providing insightful comments on the manuscript and the web application interface. I also thank Jessalyn Lu and Yin Yin Lu for proofreading the manuscript, and the user community for feedback.
Notably, the supplemental PDF has been diligently written to serve as a practical guide, with complete and reproducible examples. In fact, the paper concludes with the following two sentences:
Finally, all results reported here are completely reproduceable (see the supplemental PDF). Any questions related to this work are welcome and will be openly addressed on the 3DNA Forum (http://forum.x3dna.org).

As of version 2.0 (to be released soon), DSSR has a new module for in silico base mutations that is context sensitive. Powered by the DSSR analysis engine, the module allows users to perform base mutations in unprecedented flexibility and convenance. Here are some examples:
- Mutate all bases in hairpin loops to a specific base (e.g., G)
- Mutate all non-stem bases to a specific base (e.g., U)
- Mutate bases 2-12 to a specific base (e.g., A) regardless of context
- Mutate bases 1-10 in a given structure to a new sequence (e.g., AUAUAUAUAU)
- Mutate all bases of the same type to another (e.g., A to G)
- Mutate all bases of the same type to another (e.g., C to U) except for some nucleotides
- Mutate all G-C Watson-Crick (WC) pairs to C-G WC pairs, and A-U to U-A
- Mutate all G-tetrads in G-quadruplexes to non-G-tetrads (e.g., U-tetrads)
By default, the mutation preserves both the geometry of the sugar-phosphate backbone and the base reference frame (position and orientation). As a result, re-analyzing the mutated model gives the same base-pair and step parameters as those of the original structure.
Over the years, the 3DNA mutate bases program has been cited in the literature and patent, including the following ones:
- Howe, John A., et al. Selective small-molecule inhibition of an RNA structural element. Nature 526.7575 (2015): 672-677.
- Wang, Hao, et al. Dual-targeting small-molecule inhibitors of the Staphylococcus aureus FMN riboswitch disrupt riboflavin homeostasis in an infectious setting. Cell Chemical Biology 24.5 (2017): 576-588.
- AlQuraishi, Mohammed, and Harley H. McAdams. Three enhancements to the inference of statistical protein‐DNA potentials. Proteins: Structure, Function, and Bioinformatics 81.3 (2013): 426-442.
- Wang, Harris, Sagi Shapira, and Victoria Stockman. High-throughput strategy for dissecting mammalian genetic interactions. U.S. Patent Application No. 15/747,677.
The DSSR mutation module has completely obsoleted the mutate_bases program distributed in 3DNA v2.x. In addition to serving as a drop-in replacement of mutate_bases, the DSSR approach offers much more features and versatility: it is simply better.

I recently noticed a bioRxiv preprint, titled Role of RNA Guanine Quadruplexes in Favoring the Dimerization of SARS Unique Domain in Coronaviruses by a European team consisting of scientists from France, Italy, and Spain. The abstract is as follows. Figure 1 shows a schematic representation of the mRNA with a G-Quadruplex structure, functioning in a healthy cell and an infected cell by coronavirus.
Coronaviruses may produce severe acute respiratory syndrome (SARS). As a matter of fact, a new SARS-type virus, SARS-CoV-2, is responsible of a global pandemic in 2020 with unprecedented sanitary and economic consequences for most countries. In the present contribution we study, by all-atom equilibrium and enhanced sampling molecular dynamics simulations, the interaction between the SARS Unique Domain and RNA guanine quadruplexes, a process involved in eluding the defensive response of the host thus favoring viral infection of human cells. The results obtained evidence two stable binding modes with guanine quadruplexes, driven either by electrostatic (dimeric mode) or by dispersion (monomeric mode) interactions, are proposed being the dimeric mode the preferred one, according to the analysis of the corresponding free energy surfaces. The effect of these binding modes in stabilizing the protein dimer was also assessed, being related to its biological role in assisting SARS viruses to bypass the host protective response. This work also constitutes a first step of the possible rational design of efficient therapeutic agents aiming at perturbing the interaction between SARS Unique Domain and guanine quadruplexes, hence enhancing the host defenses against the virus.
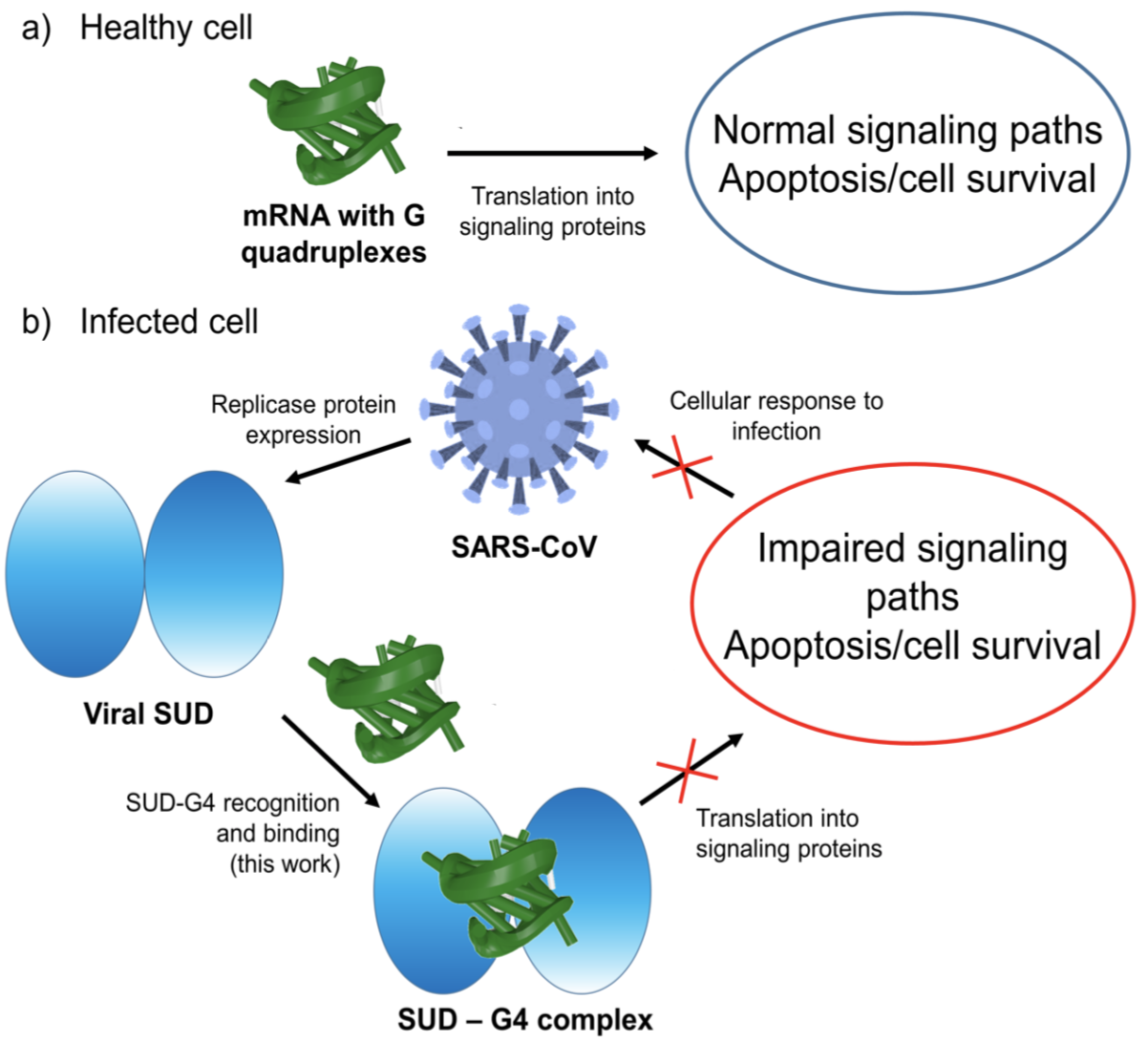
Figure 1) Schematic representation of the mRNA function in a) a healthy cell and b) an infected cell by coronavirus. Panel b) showcases the influence of viral SUD binding to G4 sequences of mRNA that encodes crucial proteins for the apoptosis/cell survival regulation and other signaling paths.
In the manuscript, the software tools employed in this MD study are described as below:
… Both protein and RNA have been described with the amber force field including the bsc1 corrections, and the MD simulations have been performed in the constant pressure and temperature ensemble (NPT) at 300K and 1 atm. All MD simulations have been performed using the NAMD code and analyzed via VMD, the G4 structure has also been analyzed with the 3DNA suite.
I am glad that 3DNA has played a role in the analysis of G-quadruplexes in this timely contribution. In particular, I would like to draw attention of the community to 3DNA-DSSR which has a brand-new module dedicated to the automatic identification and comprehensive characterization of G-quadruplexes. The DSSR-annotated G-quadruplexes from the PDB should be of great interest to a wide audience, especially the experimentalists. As a concrete example, the authors noted that “The crystal structure … of the oligonucleotide (pdb 1J8G) have been chosen coherently with the experimental work performed by Tan et al”. Follow the link to see results of DSSR-derived G-quadruplex features in PDB entry 1J8G and you are guaranteed to see features not available elsewhere.
Note added on July 9, 2020: This paper has been published in J. Phys. Chem. Lett. 2020, 11, 5661−5667.

The Kribelbauer et al. article, Towards a mechanistic understanding of DNA methylation readout by transcription factors has recently been published in the Journal of Molecular Biology (JMB). I am honored to be among the author list, and I learned a lot during the process. For the project, I added the --methyl-C (short-form: --5mc) option to SNAP (v1.0.6-2019sep30) for the automatic identification and annotation of DNA-transcription factor (TF) complexes containing 5-methyl-cytosine (5mC). The results are presented in a dynamic table, easily accessible at URL http://snap-5mc.x3dna.org, and summarized in Fig. 1 “Structural basis of how TFs recognize methylated DNA” (see below) of the JMB paper.

Details on the SNAP-enabled curation of TF-DNA complexes containing 5mC from atomic coordinates in the Protein Data Bank (PDB) are available in a tutorial page at http://snap-5mc.x3dna.org/tutorial. In essence, the process can be easily understood via a concrete example with PDB id 4m9e, as shown below.
x3dna-snap --methyl-C --type=base -i=4m9e.pdb -o=4m9e-5mC.out
Here the --methyl-C option is specific for 5mC-DNA, and --type=base ensures that at least one DNA base atom is contacting protein amino acid(s). If these conditions are fulfilled, SNAP would produce two additional 5mC-related files, apart from the normal output file (i.e., 4m9e-5mC.out, as specified in the example):
- 4m9e-5mC.txt — a simple text file with the following contents:
4m9e:B.5CM5: stacking-with-A.ARG443 is-WC-paired is-in-duplex [+]:GcG/cGC
4m9e:C.5CM5: other-contacts is-WC-paired is-in-duplex [-]:cGT/AcG
- 4m9e-5mC.pdb — a corresponding PDB file, potentially multi-model, two as in this case. Moreover, the cluster of interacting residues (DNA nucleotides and protein amino acids) is oriented in the standard base reference frame of 5mC, allowing for easy comparison and direct overlap of multiple clusters.
In practice, SNAP needs to take care of many details for the automatic identification and annotation of 5mC-DNA-TF complexes directly from PDB entries. For example, 5mC in DNA is designated 5CM and the 5-methyl carbon atom is named C5A in the PDB (see the blogpost 5CM and 5MC, two forms of 5-methylcytosine in the PDB). Moreover, the --type=base option is employed to ensure that base atoms (regardless sugar-phosphate atoms) of 5mC are directly involved in interactions with amino acids.
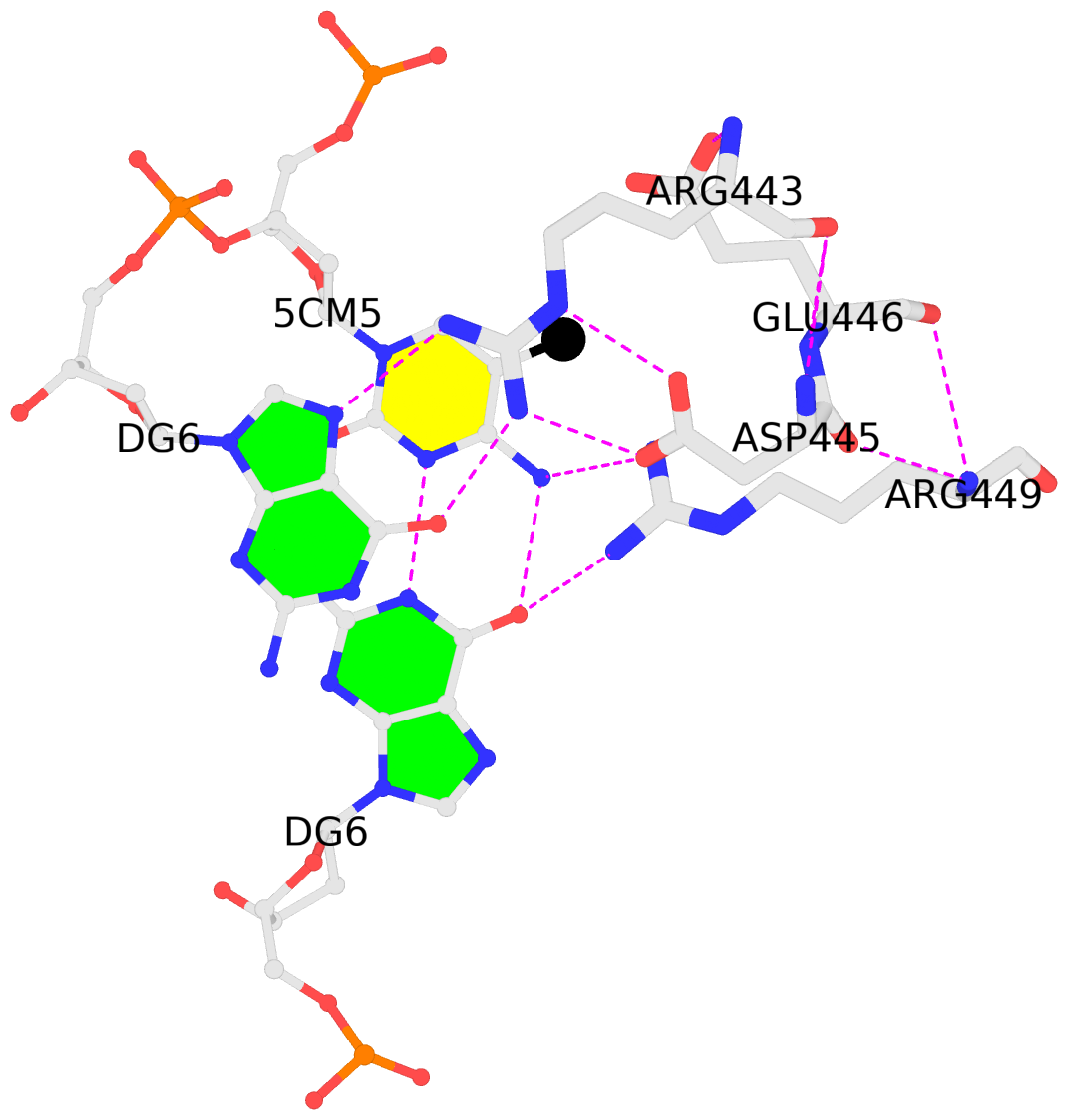
It is also worth noting the combined use of DSSR for the generation of molecular images (rendered with PyMOL), as shown below. Here the DSSR options --block-file=fill-hbond (fill to fill base rings and hbond to draw hydrogen bonds) and --cartoon-block=sticks-label are used. The 3DNA DSSR/SNAP combo is a unique and powerful toolset for structural bioinformatics, as demonstrated in DNAproDB from the Rohs lab (see my blogpost SNAP and DSSR in DNAproDB). The JMB paper represents yet another example. I can only expect to see more combined DSSR/SNAP applications in the future.

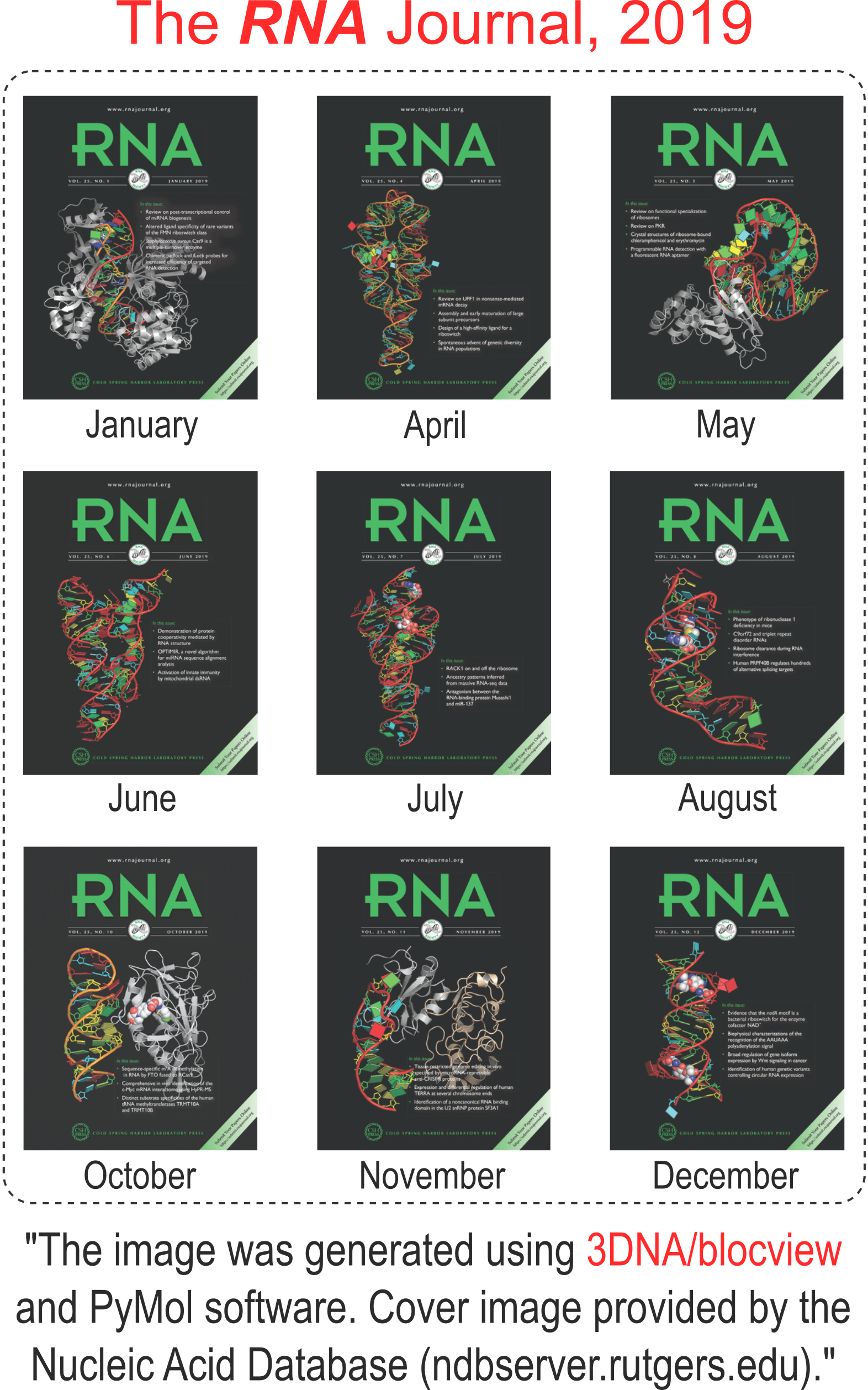
I recently performed a quick survey of the cover images of the RNA journal in 2019. I was pleased to find that 9 out of the 12 cover images were provided by the Nucleic Acid Database where 3DNA/blockview and PyMOL were employed, as noted below:
The RNA backbone is displayed as a red ribbon; bases are shown as blocks with NDB coloring: A—red, C—yellow, G—green, U—cyan; geneticin ligands are shown in spacefill with element colors: C—white, N—blue, O—red. The image was generated using 3DNA/blocview and PyMol software.
Details of the 9 cover images are listed below:
- January 2019 Rhodobacter sphaeroides Argonaute with guide RNA/target DNA duplex containing noncanonical A-G pair (PDB code: 6d9k)
- April 2019 Group I self-splicing intron P4-P6 domain mutant U131A (PDB code: 6d8l)
- May 2019 Crystal structure of T. thermophilus 50S ribosomal protein L1 in complex with helices H76, H77, and H78 of 23S RNA (PDB code: 5npm)
- June 2019 Crystal structure of ykoY-mntP riboswitch chimera bound to cadmium (PDB code: 6cc3)
- July 2019 G96A mutant of the PRPP riboswitch from T. mathranii bound to ppGpp (PDB code: 6ck4)
- August 2019 Crystal structure of the metY SAM V riboswitch (PDB code: 6fz0)
- October 2019 Crystal structure of protease factor Xa bound to RNA aptamer 11F7t and rivaroxaban (PDB code: 5vof)
- November 2019 Drosophila melanogaster nucleosome remodeling complex (PDB code: 6f4g)
- December 2019 Crystal structure of the Homo Sapiens cytoplasmic ribosomal decoding site in complex with Geneticin (PDB code: 5xz1)
Here is the composite figure of the 9 cover images.

See also:

I’ve created a web API to DSSR and SNAP, and fiber models. The overall help message is available via http://api.x3dna.org. Individually, each program is accessed as below.
Usage with 'http' (HTTPie):
http -f http://api.x3dna.org/dssr [options] url=|model@
http http://api.x3dna.org/dssr/help -- display this help message
Options:
json=true-or-FALSE(default) [e.g., json=true # JSON output]
pair=true-or-FALSE(default) [e.g., pair=1 # base-pair only]
hbond=true-or-FALSE(default) [e.g., hbond=t # H-bonding info]
more=true-or-FALSE(default) [e.g., more=y # further details]
Required parameter:
url=URL-to-coordinate-file [e.g., url=https://files.rcsb.org/download/1ehz.pdb.gz]
model@coordinate-file [e.g., model@1ehz.cif]
# Only one must be specified. 'url' precedes 'model' when both are specified.
# The coordinate file must be in PDB or PDBx/mmCIF format, optionally gzipped.
Examples:
http -f http://api.x3dna.org/dssr url=https://files.rcsb.org/download/1ehz.pdb.gz
http -f http://api.x3dna.org/dssr model@1ehz.cif pair=1
# with 'curl'
curl http://api.x3dna.org/dssr -F 'url=https://files.rcsb.org/download/1ehz.pdb.gz'
curl http://api.x3dna.org/dssr -F 'model=@1msy.pdb' -F 'pair=1'
Note:
The web API has an upper limit on coordinate file size (gzipped): < 6 MB
Usage with 'http' (HTTPie):
http -f http://api.x3dna.org/snap [options] url=|model@
http http://api.x3dna.org/snap/help -- display this help message
Options:
json=true-or-FALSE(default) [e.g., json=true # JSON output]
hbond=true-or-FALSE(default) [e.g., hbond=t # H-bonding info]
Required parameter:
url=URL-to-coordinate-file [e.g., url=https://files.rcsb.org/download/1oct.pdb.gz]
model@coordinate-file [e.g., model@1oct.cif]
# Only one must be specified. 'url' precedes 'model' when both are specified.
# The coordinate file must be in PDB or PDBx/mmCIF format, optionally gzipped.
Examples:
http -f http://api.x3dna.org/snap url=https://files.rcsb.org/download/1oct.pdb.gz
http -f http://api.x3dna.org/snap model@1oct.cif json=1
# with 'curl'
curl http://api.x3dna.org/snap -F 'url=https://files.rcsb.org/download/1oct.pdb.gz'
curl http://api.x3dna.org/snap -F 'model=@1oct.cif' -F 'json=1'
Note:
The web API has an upper limit on coordinate file size (gzipped): < 6 MB
Usage with 'http' (HTTPie):
http http://api.x3dna.org/fiber/help # display this help message
http http://api.x3dna.org/fiber/list # show a list of available fiber models (56 in total)
http http://api.x3dna.org/fiber/str_id # build model 'str_id' in the range of [1, 56]
http http://api.x3dna.org/fiber/name # generate a model with common names as shown below:
A-DNA, B-dna, C_DNA, D-DNA, ZDNA, RNA, RNAduplex, PaulingTriplex, G4
Case does not matter, and the separator can be '-' or '_' or omitted.
So a-dna, A-dNA, a_DNA, or ADNA is valid for building an A-DNA model.
Options (via query strings, or form fields):
seq=base-sequence # A, C, G, T, U for generic model
repeat=number # number of repeats of the sequence
cif=1 # output file in mmCIF format
Examples with 'http' (HTTPie):
http http://api.x3dna.org/fiber/1 # model no. 1 (i.e., calf thymus A-DNA model)
http -f http://api.x3dna.org/fiber/1 seq=A3TTT repeat=2 # specific sequence, repeated twice
http http://api.x3dna.org/fiber/rna # single-stranded RNA model
http http://api.x3dna.org/fiber/rna-ds # double-stranded RNA model
http http://api.x3dna.org/fiber/pauling # the triplex model of Pauling & Corey
http http://api.x3dna.org/fiber/g4 # G-quadruplex model
# with 'curl'
curl http://api.x3dna.org/fiber/1
curl http://api.x3dna.org/fiber/1 -d 'seq=A3TTT' -d 'repeat=2'
curl http://api.x3dna.org/fiber/rna
curl http://api.x3dna.org/fiber/rna-ds
curl http://api.x3dna.org/fiber/pauling
curl http://api.x3dna.org/fiber/g4
Note:
The web API has two upper limits: repeats < 1,000, and nucleotides < 10,000.

The skmatic.x3dna.org website (see screenshot below) aims to showcase DSSR-enabled cartoon-block schematics of nucleic acid structures using PyMOL. It presents a simple interface to browse pre-calculated PDB entries with a set of default settings: long rectangular blocks for Watson-Crick base-pairs, square blocks for G-tetrads in G-quadruplexes, with minor-groove edges in black. Users can also specify an URL to a PDB- or mmCIF-formatted file or upload such an atomic coordinates file directly, and set several common options to customerize to the rendered image.
Moreover, a web API to DSSR-PyMOL schematics is available to allow for its easy integration into third-party tools.

Input a PDB id
Pre-calculated cartoon-block images together with summary information are available for PDB entries of nucleic-acid-containing structures. Note that gigantic structures like ribosomes that are only represented in mmCIF format are excluded from the resource. The base block images are most effective for small to medium-sized structures.
Here are a few examples:
- 1ehz, the crystal structure of yeast phenylalanine tRNA at 1.93-Å resolution
- 2lx1, the major conformation of the internal loop 5’GAGU/3’UGAG
- 2grb”, the crystal structure of an RNA quadruplex containing inosine-tetrad
- 4da3, the crystal structure of an intramolecular human telomeric DNA G-quadruplex 21-mer bound by the naphthalene diimide compound MM41
- 1oct, crystal structure of the Oct-1 POU domain bound to an octamer site
- 2hoj, the crystal structure of an E. coli thi-box riboswitch bound to thiamine pyrophosphate, manganese ions
Each entry is shown with images in six orthogonal perspectives: front, back, right, left, top, bottom. The ‘front’ image (upper-left in the panel) is oriented into the most-extended view with the DSSR --blocview option. The corresponding PyMOL session file and PDB coordinate file are available for download. One can also visualize the structure interactively via 3Dmol.js.
Sample PDB entries
Users can browse random samples of pre-calculated PDB entries. The number should be between 3 and 99, with a default of 12 entries (see below for an example). Simply click the ‘Submit’ button or the “Random samples (3 to 99)”: http://skmatic.x3dna.org/pdb_entry link to see results of randomly picked 12 PDB entries each time.
Specify a coordinate file
The atomic coordinate file must be in PDB or mmCIF format, and can be optionally gzipped (.gz). One can either specify an URL to or select a coordinate file. Several common options are available to allow for user customizations.
Web API help message
Usage with 'http' (HTTPie):
http -f http://skmatic.x3dna.org/api [options] url=|model@
http http://skmatic.x3dna.org/api/pdb/pdb_id -- for a pre-calculated PDB entry
http http://skmatic.x3dna.org/api/help -- display this help message
Options:
block_file=styles-in-free-text-format [e.g., block_file=wc-minor]
block_color=nt-selection-and-color [e.g., block_color='A:pink']
block_depth=thickness-of-base-block [e.g., block_depth=1.2]
r3d_file=true-or-FALSE(default) [e.g., r3d_file=true]
raw_xyz=true-or-FALSE(default) [e.g., raw_xyz=true]
Required parameter
url=URL-to-coordinate-file [e.g., url=https://files.rcsb.org/download/1ehz.pdb.gz]
model@coordinate-file [e.g., model@1ehz.cif]
# Only one must be specified. 'url' precedes 'model' when both are specified.
# The coordinate file must be in PDB or PDBx/mmCIF format, optionally gzipped.
Examples
http -f http://skmatic.x3dna.org/api block_file='wc-minor' model@1ehz.cif r3d_file=t
http -f http://skmatic.x3dna.org/api url=https://files.rcsb.org/download/1ehz.pdb.gz -d -o 1ehz.png
http http://skmatic.x3dna.org/api/pdb/1ehz -d -o 1ehz.png
# with 'curl'
curl http://skmatic.x3dna.org/api -F 'model=@1msy.pdb' -F 'block_file=wc-minor' -F 'r3d_file=1'
curl http://skmatic.x3dna.org/api -F 'url=https://files.rcsb.org/download/1ehz.pdb.gz' -o 1ehz.png
curl http://skmatic.x3dna.org/api/pdb/1ehz -o 1ehz.png
Sample images

While reading DNAproDB: an expanded database and web-based tool for structural analysis of DNA–protein complexes, I noticed SNAP and DSSR being mentioned. The detailed citations are as below:
Information about individual nucleotide–residue interactions is also provided, such as hydrogen bonding, interaction geometry (based on SNAP (10)), buried solvent accessible surface areas and identification of the interacting residue and nucleotide moieties …
DNAproDB assigns a geometry for every nucleotide–residue interaction identified using SNAP, a component of the 3DNA program suite (10). The residues for which probabilities are shown are those with planar side chains so that a stacking conformation can be defined.
Base pairing and base stacking between nucleotides are identified using the program DSSR (20).
SNAP and DSSR are two (relatively) new programs in the 3DNA software suite. As the author, I am always glad to see them being cited explicitly in literature. The fact that SNAP and DSSR are cited together by DNAproDB, however, is especially significant. I am aware of the initial version of DNAproDB, but I definitely like the updated one better. This is what I recently wrote in response to a question on the 3DNA Forum:
Regarding DNA-protein interactions in general, you may want to have a look of DNAproDB from the Remo Rohs laboratory. A new paper has just been published in NAR, ‘DNAproDB: an expanded database and web-based tool for structural analysis of DNA–protein complexes’.
I’ve no doubt that SNAP and DSSR would be widely used in applications related to DNA/RNA structural bioinformatics. DSSR (to a lesser extent, SNAP) represents my view of what a scientific software tool should be.

Recently I noticed two new citations to DSSR, an integrated software tool for dissecting the spatial structure of RNA. One is from the Yesselman et al. article Computational design of three-dimensional RNA structure and function in Nature Nanotechnology, and the other is from the Wang et al. article 3dRNA v2.0: An Updated Web Server for RNA 3D Structure Prediction in International Journal of Molecular Sciences.
Yesselman et al. has used DSSR in RNAMake for building the motif library. The relevant section is as follows:
We processed each RNA structure to extract every motif with Dissecting the Spatial Structure of RNA (DSSR)54 with the following command:
x3dna-dssr –i file.pdb –o file_dssr.out
We manually checked each extracted motif to confirm that it was the correct type, as DSSR sometimes classifies tertiary contacts as higher-order junctions and vice versa. For each motif collected from DSSR, we ran the X3DNA find_pair and analyze programs to determine the reference frame for the first and last base pair of each motif to allow for the alignment between motifs:
find_pair file.pdb 2> /dev/null stdout | analyze stdin >& /dev/null
It is worth noting the sentence that “DSSR sometimes classifies tertiary contacts as higher-order junctions and vice versa.” Presumably. the authors are referring to the inclusion of ‘isolated canonical pairs’ in junctions by default in DSSR. Overall, the default DSSR settings follow the most common practice in RNA literature. In the meantime, I am aware that the community may not agree on every detail. Thus DSSR provide many options (documented or otherwise) to cater for other potential use cases. See the Stems of junction structure have only one base pair and Junction definition threads on the 3DNA Forum for two examples. In the long run, DSSR is likely to help consolidate RNA nomenclature that can be applied in a pragmatic, consistent manner.
Note also that DSSR provides the reference frame of each identified base pair via the JSON option. Using 1ehz as an example, the following command provides detailed information about base pairs:
x3dna-dssr -i=1ehz.pdb --json --more | jq .pairs
In the 3dRNA 2.0 paper, DSSR is cited as below. This is the first time DSSR is integrated in the 3dRNA pipeline.
The predicted structures are built from the sequence and secondary structure, while the former is obtained from their native structures fetched from PDB (https://www.rcsb.org/), and the latter is calculated from DSSR (Dissecting the Spatial Structure of RNA) [39].